seananderson / Glmmfields
Programming Languages
Labels
Projects that are alternatives of or similar to Glmmfields
glmmfields 
The glmmfields R package implements Bayesian spatiotemporal models that allow for extreme spatial deviations through time. It uses a predictive process approach with random fields implemented through a multivariate-t distribution instead of a multivariate normal. The models are fit with Stan.
We published a paper describing the model and package in Ecology:
Anderson, S. C., Ward, E. J. 2019. Black swans in space: modelling spatiotemporal processes with extremes. 100(1):e02403. https://doi.org/10.1002/ecy.2403
You can install the CRAN version of the package with:
install.packages("glmmfields")
If you have a C++ compiler installed, you can install the development version of the package with:
# install.packages("devtools")
devtools::install_github("seananderson/glmmfields", build_vignettes = TRUE)
glmmfields can also fit spatial GLMs with Stan. See the vignette:
vignette("spatial-glms", package = "glmmfields")
An example spatiotemporal model
library(glmmfields)
#> Loading required package: Rcpp
library(ggplot2)
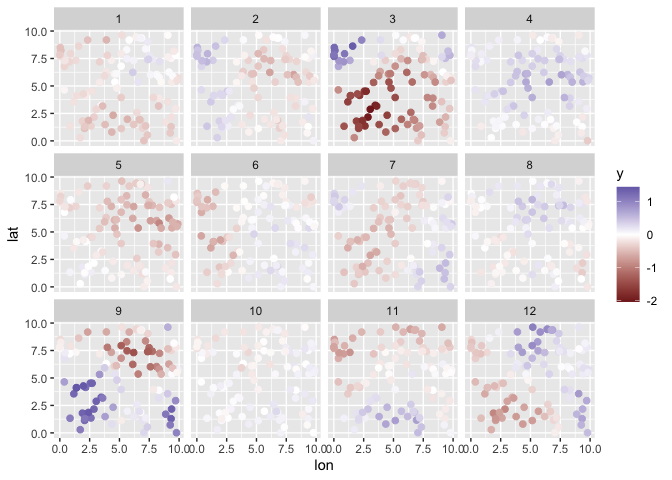
Simulate data:
set.seed(42)
s <- sim_glmmfields(
df = 2.8, n_draws = 12, n_knots = 12, gp_theta = 2.5,
gp_sigma = 0.2, sd_obs = 0.1
)
head(s$dat)
#> time pt y lon lat station_id
#> 1 1 1 0.02818963 9.148060 6.262453 1
#> 2 1 2 -0.21924739 9.370754 2.171577 2
#> 3 1 3 -0.34719485 2.861395 2.165673 3
#> 4 1 4 -0.15785483 8.304476 3.889450 4
#> 5 1 5 -0.04703617 6.417455 9.424557 5
#> 6 1 6 -0.23904924 5.190959 9.626080 6
print(s$plot)
Fit the model:
options(mc.cores = parallel::detectCores()) # for parallel processing
m <- glmmfields(y ~ 0,
data = s$dat, time = "time",
lat = "lat", lon = "lon",
nknots = 12, estimate_df = TRUE, iter = 800, seed = 1
)
print(m)
#> Inference for Stan model: glmmfields.
#> 4 chains, each with iter=800; warmup=400; thin=1;
#> post-warmup draws per chain=400, total post-warmup draws=1600.
#>
#> mean se_mean sd 2.5% 25% 50% 75% 97.5% n_eff Rhat
#> df[1] 3.78 0.04 1.40 2.09 2.73 3.45 4.44 7.46 1422 1.00
#> gp_sigma 0.30 0.00 0.04 0.22 0.27 0.30 0.33 0.39 441 1.01
#> gp_theta 2.59 0.00 0.06 2.46 2.55 2.59 2.63 2.71 1493 1.00
#> sigma[1] 0.10 0.00 0.00 0.09 0.10 0.10 0.10 0.10 1695 1.00
#> lp__ 2291.18 0.41 9.24 2271.74 2285.06 2291.84 2297.72 2307.86 516 1.01
#>
#> Samples were drawn using NUTS(diag_e) at Sat Jan 9 09:30:08 2021.
#> For each parameter, n_eff is a crude measure of effective sample size,
#> and Rhat is the potential scale reduction factor on split chains (at
#> convergence, Rhat=1).
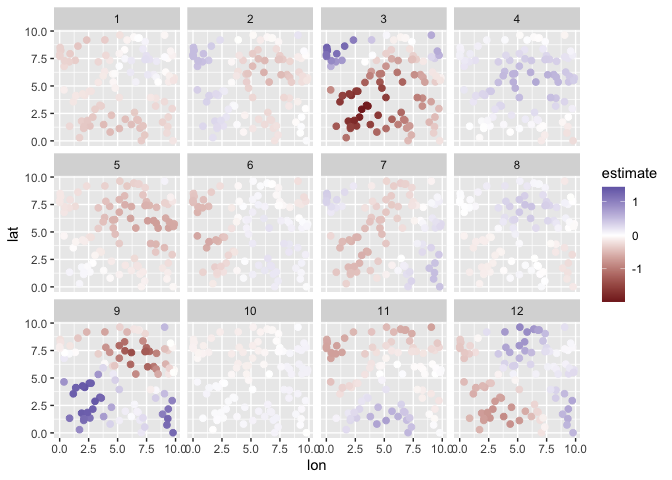
Plot:
plot(m, type = "prediction") + scale_color_gradient2()
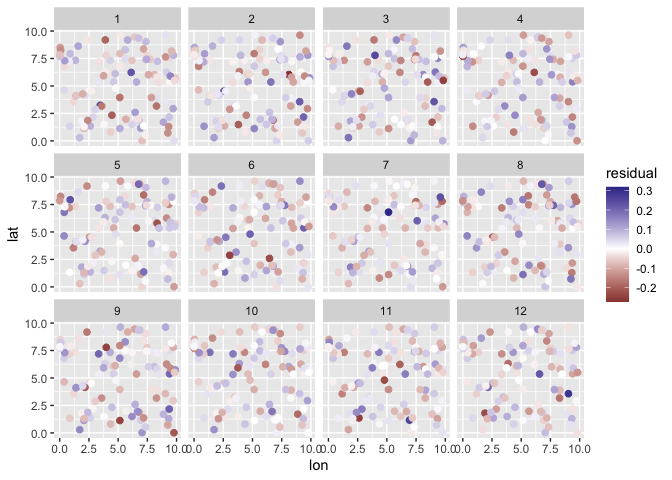
plot(m, type = "spatial-residual")
Predictions:
# link scale:
p <- predict(m)
head(p)
#> # A tibble: 6 x 3
#> estimate conf_low conf_high
#> <dbl> <dbl> <dbl>
#> 1 -0.0292 -0.0864 0.0312
#> 2 -0.291 -0.361 -0.219
#> 3 -0.397 -0.451 -0.344
#> 4 -0.194 -0.271 -0.120
#> 5 -0.0367 -0.111 0.0416
#> 6 -0.216 -0.294 -0.140
# posterior predictive intervals on new observations (include observation error):
p <- predictive_interval(m)
head(p)
#> # A tibble: 6 x 3
#> estimate conf_low conf_high
#> <dbl> <dbl> <dbl>
#> 1 -0.0292 -0.237 0.178
#> 2 -0.291 -0.503 -0.0869
#> 3 -0.397 -0.603 -0.195
#> 4 -0.194 -0.401 0.0137
#> 5 -0.0367 -0.242 0.172
#> 6 -0.216 -0.425 -0.00916
Use the tidy method to extract parameter estimates as a data frame:
x <- tidy(m, conf.int = TRUE)
#> Registered S3 method overwritten by 'broom.mixed':
#> method from
#> tidy.gamlss broom
head(x)
#> # A tibble: 6 x 5
#> term estimate std.error conf.low conf.high
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 df[1] 3.45 1.40 2.09 7.46
#> 2 gp_sigma 0.299 0.0442 0.217 0.389
#> 3 gp_theta 2.59 0.0640 2.46 2.71
#> 4 sigma[1] 0.0979 0.00216 0.0941 0.102
#> 5 spatialEffectsKnots[1,1] -0.110 0.0355 -0.182 -0.0413
#> 6 spatialEffectsKnots[2,1] -0.230 0.0357 -0.302 -0.163
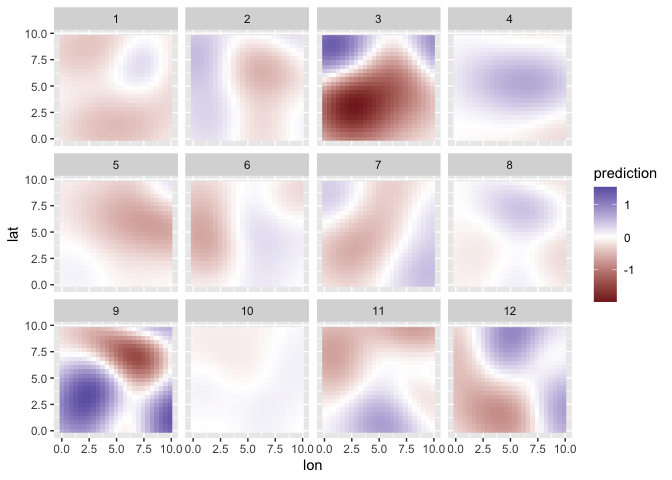
Make predictions on a fine-scale spatial grid:
pred_grid <- expand.grid(
lat = seq(min(s$dat$lat), max(s$dat$lat), length.out = 25),
lon = seq(min(s$dat$lon), max(s$dat$lon), length.out = 25),
time = unique(s$dat$time)
)
pred_grid$prediction <- predict(m,
newdata = pred_grid, type = "response", iter = 100, estimate_method = "median"
)$estimate
ggplot(pred_grid, aes(lon, lat, fill = prediction)) +
facet_wrap(~time) +
geom_raster() +
scale_fill_gradient2()
References
Anderson, S. C., Ward, E. J. 2019. Black swans in space: modelling spatiotemporal processes with extremes. 100(1):e02403. https://doi.org/10.1002/ecy.2403
Latimer, A. M., S. Banerjee, H. Sang Jr, E. S. Mosher, and J. A. Silander Jr. 2009. Hierarchical models facilitate spatial analysis of large data sets: a case study on invasive plant species in the northeastern United States. Ecology Letters 12:144–154.
Shelton, A. O., J. T. Thorson, E. J. Ward, and B. E. Feist. 2014. Spatial semiparametric models improve estimates of species abundance and distribution. Canadian Journal of Fisheries and Aquatic Sciences 71:1655–1666.