Urinx / Alphafold_pytorch
Programming Languages
Labels
AlphaFold - PyTorch
This project provides an implementation of the DeepMind's AlphaFold based on PyTorch for research, also includes the converted model weights and inputs. Note that this code can also works well on the original .ckpt format model weights and .tfrec format inputs.
The original DeepMind's implementation is based on TensorFlow, related publication paper is Senior, A.W., Evans, R., Jumper, J. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020). [code] [paper]
Comparison
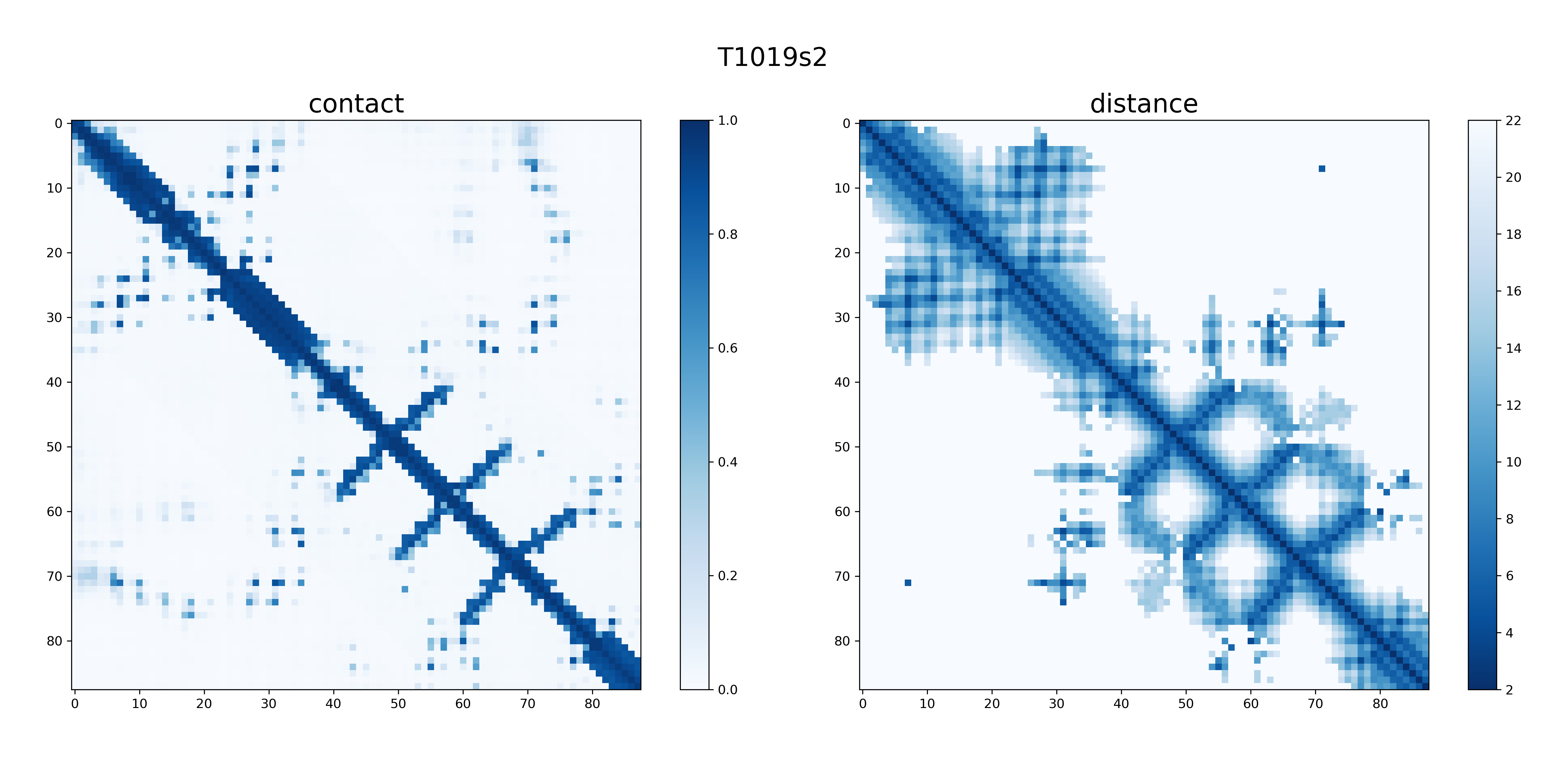
I calculated the differences of the final output distogram probs between the PyTorch version and original TensorFlow version. The distogram probs is full set of distance distribution predictions constructed by combining such predictions that covers the entire distance map, which size is LxLx64. Take target T1019s2 for example, the error of distogram probs (88x88x64) between these two results is 0.467 per channel. The picture above is the result of T1019s2.
Despite the speed is around 7 times slower than the TensorFlow version, I still recommend this project for you because I refactored the entire code and provided a simplest way for you to understand AlphaFold. In addition, contrast to TensorFlow, PyTorch is imperative and you can simply throw in a pdb breakpoint anywhere into your model.
Usage
Dependencies
- Python 3.6+
- PyTorch 1.3+
- TensorFlow 2.0+ (This is optional if you want to load original
.ckptformat model weights and.tfrecformat inputs)
Run example prediction
You can use the alphafold.sh script to run the entire Distogram prediction system.
./alphafold.sh
You can simply modify the keywords such as TARGET, TARGET_FILE in this file to run prediction for other targets.
Detailed script usage
> python alphafold.py -h
usage: alphafold.py [-h] -i INPUT [-o OUT] [-m MODEL] [-r REPLICA]
[-t {D,B,T}] [-e]
optional arguments:
-h, --help show this help message and exit
-i INPUT, --input INPUT
target protein, support both .pkl or .tfrec format
-o OUT, --out OUT output dir
-m MODEL, --model MODEL
model dir
-r REPLICA, --replica REPLICA
model replica
-t {D,B,T}, --type {D,B,T}
model type: D - Distogram, B - Background, T - Torsion
-e, --ensemble ensembling all replica outputs
For example:
python alphafold.py -i test_data/T1019s2.pkl -o T1019s2_out -t D -r 0
This uses the replica 0 of Distogram models to predict the distogram probs of the input data.
It also supports tensorflow data input and model:
python alphafold.py -i test_data/T1019s2.tfrec -o T1019s2_out -m tf_model_path/
Data
Model weights
All converted model weights data can be downloaded from http://bit.ly/alphafold-model. The weights data are in a zip file which has about 210 MB, in the model folder which contains:
- A directory
873731. This contains the weights for the distogram model. - A directory
916425. This contains the weights for the background distogram model. - A directory
941521. This contains the weights for the torsion model.
Each directory with model weights contains a number of different model configurations. Each model has a config file and associated weights. There is only one torsion model. Each model directory also contains a stats file that is used for feature normalization specific to that model.
The original TensorFlow model checkpoints can be downloaded from http://bit.ly/alphafold-casp13-weights.
Input data
For now the input data is .pkl, .npy or .tfrec format file which contains required features. The details of those feature generation can be found in the README of DeepMind's AlphaFold project.
For convenience, I provided a shell script feature.sh to generate those required features data from given target sequence (.seq file, fasta format). Before run this script, there are a few steps you need to start with:
- Setup PSI-BLAST from NCBI BLAST.
- Setup HHBlits from HH-suite3.
# Installation HH-suite git clone https://github.com/soedinglab/hh-suite.git mkdir -p hh-suite/build && cd hh-suite/build cmake -DCMAKE_INSTALL_PREFIX=. .. make -j 4 && make install export PATH="$(pwd)/bin:$(pwd)/scripts:$PATH" # Download Databases cd ..; mkdir databases; cd databases wget http://wwwuser.gwdg.de/~compbiol/uniclust/2018_08/uniclust30_2018_08_hhsuite.tar.gz tar xzvf uniclust30_2018_08_hhsuite.tar.gz
- Setup plmDCA, which need
MatlaborOctaveto run this code. Here I provided a modifiedplmDCA.mfile for Octave which can save the intermediate data that alphafold needs, but I haven't test it in matlab.git clone https://github.com/magnusekeberg/plmDCA.git mv plmDCA/plmDCA_asymmetric_v2 plmDCA/plmDCA cp plmDCA.m plmDCA/plmDCA_asymmetric_v2/ # mex .c file, if you use matlab you need do this in matlab console cd plmDCA/plmDCA_asymmetric_v2/functions/; for i in *.c; do octave --eval "mex $i";done cd ../3rd_party_code/minFunc/; for i in *.c; do octave --eval "mex $i"; done
- In
feature.shset the following:-
TARGETto the name of the target. -
TARGET_DIRto the path to the directory with the target input data. -
TARGET_SEQto the path of the target input seq file. -
PLMDCA_DIRto the path of plmDCA folder.
-
The example target T1019s2 input data, output results by two version AlphaFold for comparison and its generated features you get download from http://bit.ly/alphafold-T1019s2, which has about 210 MB.
The dataset to reproduce AlphaFold's CASP13 results can be downloaded from http://bit.ly/alphafold-casp13-data, which has about 43.5 GB.
Note that, profile_with_prior and profile_with_prior_without_gaps two features I can't figure it out, so it just be set to all zeros for now. Please let me know if you have any idea.