jrzech / Reproduce Chexnet
Projects that are alternatives of or similar to Reproduce Chexnet
reproduce-chexnet
Provides Python code to reproduce model training, predictions, and heatmaps from the CheXNet paper that predicted 14 common diagnoses using convolutional neural networks in over 100,000 NIH chest x-rays.
Getting Started:
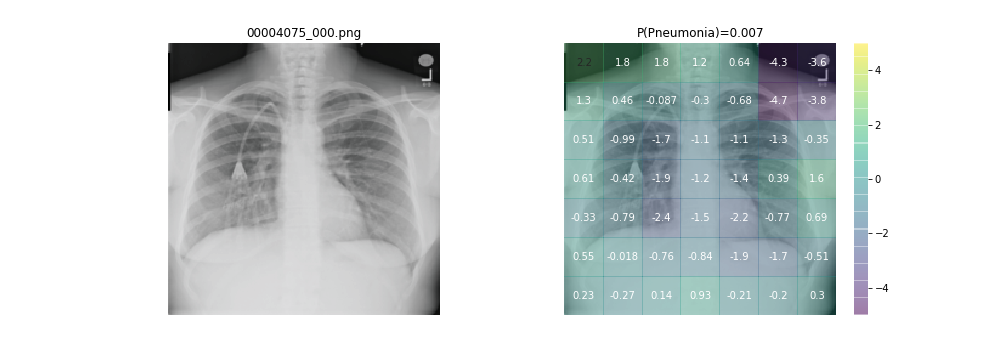
Click on the launch binder button at the top of this README to launch a remote instance in your browser using binder. This requires no local configuration and lets you get started immediately. Open Explore_Predictions.ipynb, run all cells, and follow the instructions provided to review a selection of included chest x-rays from NIH.
To configure your own local instance (assumes Anaconda is installed; can be run on Amazon EC2 p2.xlarge instance if you do not have a GPU):
cd reproduce-chexnet
conda env create -f environment.yml
source postBuild
source activate reproduce-chexnet
Replicated results:
This reproduction achieved average test set AUC 0.836 across 14 findings compared to 0.841 reported in original paper:
| retrained auc | chexnet auc | |
|---|---|---|
| label | ||
| Atelectasis | 0.8161 | 0.8094 |
| Cardiomegaly | 0.9105 | 0.9248 |
| Consolidation | 0.8008 | 0.7901 |
| Edema | 0.8979 | 0.8878 |
| Effusion | 0.8839 | 0.8638 |
| Emphysema | 0.9227 | 0.9371 |
| Fibrosis | 0.8293 | 0.8047 |
| Hernia | 0.9010 | 0.9164 |
| Infiltration | 0.7077 | 0.7345 |
| Mass | 0.8308 | 0.8676 |
| Nodule | 0.7748 | 0.7802 |
| Pleural_Thickening | 0.7860 | 0.8062 |
| Pneumonia | 0.7651 | 0.7680 |
| Pneumothorax | 0.8739 | 0.8887 |
Results available in pretrained folder:
-
aucs.csv: test AUCs of retrained model vs original ChexNet reported results -
checkpoint: saved model checkpoint -
log_train: log of train and val loss by epoch -
preds.csv: individual probabilities for each finding in each test set image predicted by retrained model
NIH Dataset
To explore the full dataset, download images from NIH (large, ~40gb compressed),
extract all tar.gz files to a single folder, and provide path as needed in code.
Train your own model!
Please note: a GPU is required to train the model. You will encounter errors if you do not have a GPU available and CUDA installed and you attempt to retrain. With a GPU, you can retrain the model with retrain.py. Make sure you download the full NIH dataset before trying this. If you run out of GPU memory, reduce BATCH_SIZE from its default setting of 16.
If you do not have a GPU, but wish to retrain the model yourself to verify performance, you can replicate the model using Amazon EC2's p2.xlarge instance ($0.90/hr at time of writing) with an AMI that has CUDA installed (e.g. Deep Learning AMI (Ubuntu) Version 8.0 - ami-dff741a0). After creating and ssh-ing into the EC2 instance, follow the instructions in Getting Started above to configure your environment. If you have no experience with Amazon EC2, fast.ai's tutorial is a good place to start
Note on training
I use SGD+momentum rather than the Adam optimizer as described in the original paper. I achieved better results with SGD+momentum, as has been reported in other work.
Note on data
A sample of 621 test NIH chest x-rays enriched for positive pathology is included with the repo to faciliate immediate use and exploration in the Explore Predictions.ipynb notebook. The full NIH dataset is required for model retraining.
Use and citation
My goal in releasing this code is to increase transparency and replicability of deep learning models in radiology. I encourage you to use this code to start your own projects. If you do, please cite the repo:
author = {Zech, J.},
title = {reproduce-chexnet},
year = {2018},
publisher = {GitHub},
journal = {GitHub repository},
howpublished = {\url{https://github.com/jrzech/reproduce-chexnet}}
}
Acknowledgements
With deep gratitude to researchers and developers at PyTorch, NIH, Stanford, and Project Jupyter, on whose generous work this project relies. With special thanks to Sasank Chilamkurthy, whose demonstration code was incorporated into this project. PyTorch is an incredible contribution to the research community.