theislab / Paga
Licence: bsd-3-clause
Mapping out the coarse-grained connectivity structures of complex manifolds.
Stars: ✭ 113
Labels
Projects that are alternatives of or similar to Paga
Python Projects For Beginners
Source Code for 'Python Projects for Beginners' by Connor Milliken
Stars: ✭ 111 (-1.77%)
Mutual labels: jupyter-notebook
Kerasobjectdetector
Keras Object Detection API with YOLK project 🍳
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Loandefault Prediction
Lending Club Loan data analysis
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Introspective
Repo for the ML_Insights python package
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Pytorch Generative
Easy generative modeling in PyTorch.
Stars: ✭ 112 (-0.88%)
Mutual labels: jupyter-notebook
Jupytergraffiti
Create interactive screencasts inside Jupyter Notebook that anybody can play back
Stars: ✭ 114 (+0.88%)
Mutual labels: jupyter-notebook
Pedestrian Cam
Monitoring Foot Traffic over IP Webcams with ML
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Tensorflow Nlp
NLP and Text Generation Experiments in TensorFlow 2.x / 1.x
Stars: ✭ 1,487 (+1215.93%)
Mutual labels: jupyter-notebook
Mmaml Classification
An official PyTorch implementation of “Multimodal Model-Agnostic Meta-Learning via Task-Aware Modulation” (NeurIPS 2019) by Risto Vuorio*, Shao-Hua Sun*, Hexiang Hu, and Joseph J. Lim
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Kaggle Houseprices
Kaggle Kernel for House Prices competition https://www.kaggle.com/massquantity/all-you-need-is-pca-lb-0-11421-top-4
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
V2ray Deep Packet Inspection
Notebook demo V2Ray traffic classification by deep packet inspection
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Mantranet
ManTra-Net: Manipulation Tracing Network For Detection And Localization of Image Forgeries With Anomalous Features
Stars: ✭ 114 (+0.88%)
Mutual labels: jupyter-notebook
Machine learning
参考了西瓜书,sklearn源码,李航统计学,机器学习实战、机器学习中的数学
Stars: ✭ 112 (-0.88%)
Mutual labels: jupyter-notebook
Pythondata
repo for code published on pythondata.com
Stars: ✭ 113 (+0%)
Mutual labels: jupyter-notebook
Course Content
NMA Computational Neuroscience course
Stars: ✭ 2,082 (+1742.48%)
Mutual labels: jupyter-notebook
PAGA - partition-based graph abstraction
Mapping out the coarse-grained connectivity structures of complex manifolds (Genome Biology, 2019).
PAGA is available within Scanpy through: tl.paga | pl.paga | pl.paga_path | pl.paga_compare.
Below you find links to all central example notebooks, which also allow reproducing all main figures of the paper. If you start working with PAGA, go through blood/paul15.
| notebook | system | details | reference | figure |
|---|---|---|---|---|
| blood/simulated | hematopoiesis | simulated | Krumsiek et al., Plos One (2011) | 2a |
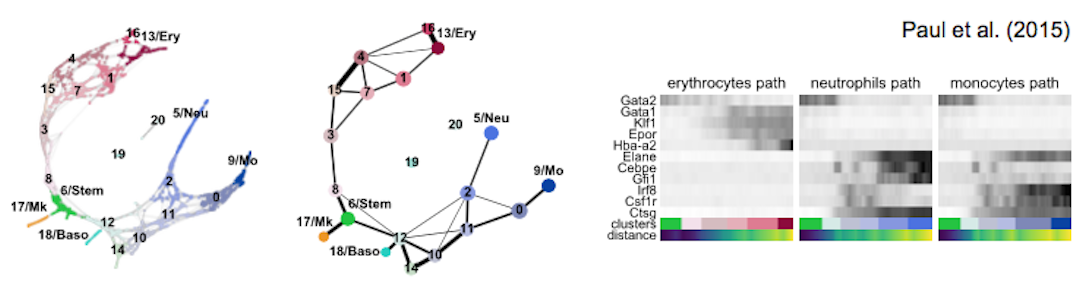
| blood/paul15 | murine hematopoiesis | 2,730 cells, MARS-seq | Paul et al., Cell (2015) | 2b |
| blood/nestorowa16 | murine hematopoiesis | 1,654 cells, Smart-seq2 | Nestorowa et al., Blood (2016) | 2c |
| blood/dahlin18 | murine hematopoiesis | 44,802 cells, 10x Genomics | Dahlin et al., Blood (2018) | 2d |
| planaria | planaria | 21,612 cells | Plass et al., Science (2018) | 3 |
| zebrafish | zebrafish embryo | 53,181 cells | Wagner et al., Science (2018) | 4 |
| 1M_neurons | neurons | 1.3 million cells, 10x Genomics | 10x Genomics (2017) | S12 |
| deep_learning | cycling Jurkat cells | 30,000 single-cell images | Eulenberg et al., Nat. Commun. (2017) | S14 |
All supplemental figures of the paper can be reproduced based on the following table.
| notebook | description | figure |
|---|---|---|
| connectivity_measure | connectivity measure | S1, S2, S3 |
| robustness | robustness and multi-resolution capacity | S4, S5 |
| comparisons/simulated_data | comparisons for simulated data | S6, S7 |
| comparisons/paul15_monocle2 | comparison Monocle 2 for Paul et al. (2015) | S8 |
| comparisons/nestorowa16_monocle2 | comparison Monocle 2 for Nestorowa et al. (2016) | S9 |
| embedding_quality | quantifying embedding quality | S10 |
| simulation | simulating hematopoiesis | S11 |
| 1M_neurons | neurons, 1.3 million cells, 10x Genomics, 10x Genomics (2017) | S12 |
| blood/paul15 | annotation of louvain clusters using PAGA | S13 |
| deep_learning | cycling Jurkat cells, 30,000 single-cell images, Eulenberg et al., Nat. Commun. (2017) | S14 |
Note that the project description data, including the texts, logos, images, and/or trademarks,
for each open source project belongs to its rightful owner.
If you wish to add or remove any projects, please contact us at [email protected].