greenelab / Tybalt
Programming Languages
Labels
Projects that are alternatives of or similar to Tybalt
Tybalt 😼
A Variational Autoencoder trained on Pan-Cancer Gene Expression
Gregory Way and Casey Greene 2017
The repository stores scripts to train, evaluate, and extract knowledge from a variational autoencoder (VAE) trained on 33 different cancer-types from The Cancer Genome Atlas (TCGA).
The specific VAE model is named Tybalt after an instigative, cat-like character in Shakespeare's "Romeo and Juliet". Just as the character Tybalt sets off the series of events in the play, the model Tybalt begins the foray of VAE manifold learning in transcriptomics. Also, deep unsupervised learning likes cats.
We discuss the training and evaluation of Tybalt in our PSB paper:
Citation
Way, GP, Greene, CS. 2018. Extracting a biologically relevant latent space from cancer transcriptomes with variational autoencoders. Pacific Symposium on Biocomputing 23:80-91. doi:10.1142/9789813235533_0008
Notes
As discovered by @enricoferrero, the preprint text (
section 2.2) states that the top median absolute deviation (MAD) genes were selected for subsetting, when the data processing code (process_data.ipynb) actually outputs the top mean absolute deviation genes. We discuss this discrepancy and its potential impact in issue #99.
git-lfs (https://git-lfs.github.com/) must be installed prior to cloning the repository. If it is not installed, run
git lfs install
The Data
TCGA has collected numerous different genomic measurements from over 10,000 different tumors spanning 33 different cancer-types. In this repository, we extract cancer signatures from gene expression data (RNA-seq).
The RNA-seq data serves as a measurement describing the high-dimensional state of each tumor. As a highly heterogeneous disease, cancer exists in several different combination of states. Our goal is to extract these different states using high capacity models capable of identifying common signatures in gene expression data across different cancer-types.
The Model
We present a variational autoencoder (VAE) applied to cancer gene expression data. A VAE is a deep generative model introduced by Kingma and Welling in 2013. The model has two direct benefits of modeling cancer gene expression data.
- Automatically engineer non-linear features
- Learning the reduced dimension manifold of cancer expression space
As a generative model, the reduced dimension features can be sampled from to simulate data. The manifold can also be interpolated to interrogate trajectories and transitions between states.
VAEs have typically been applied to image data and have demonstrated remarkable generative capacity and modeling flexibility. VAEs are different from deterministic autoencoders because of the added constraint of normally distributed feature activations per sample. This constraint not only regularizes the model, but also provides the interpretable manifold.
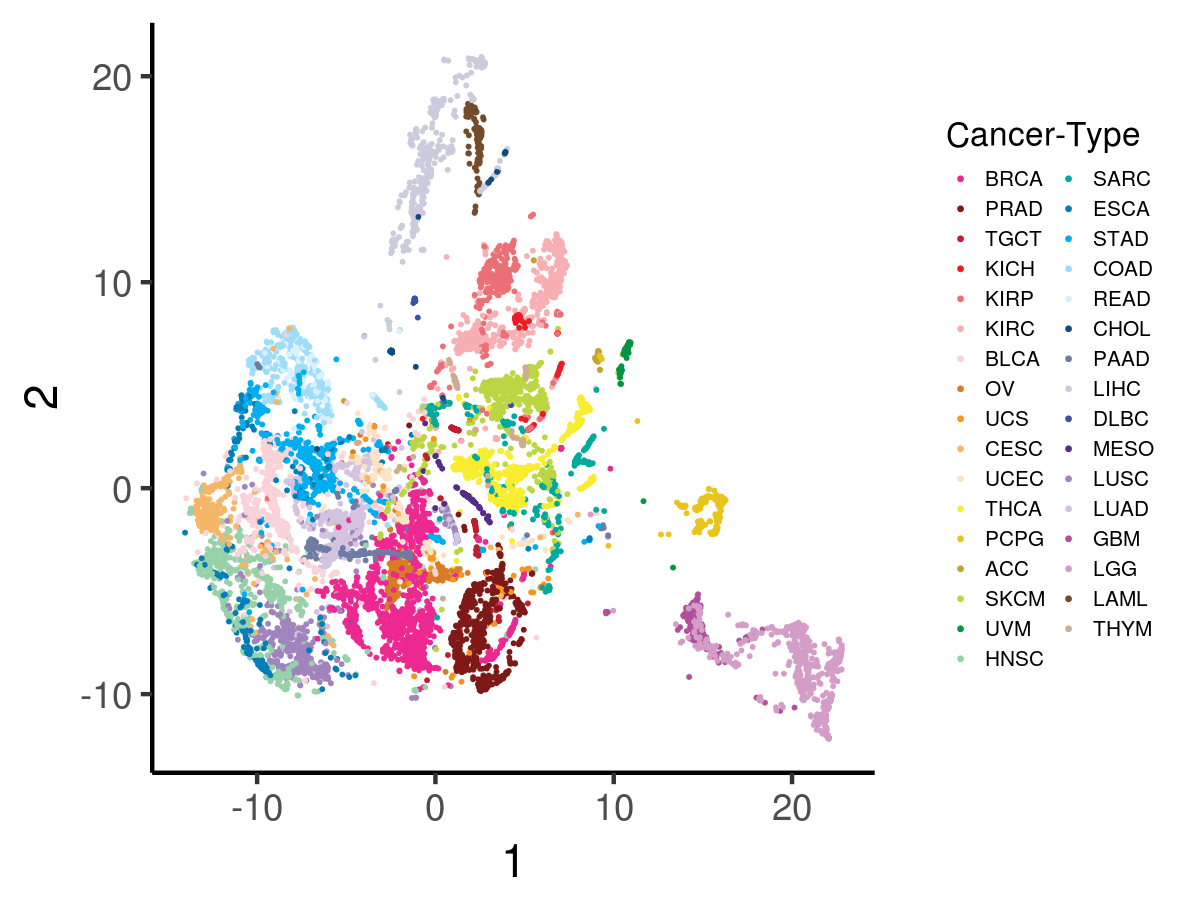
Below is a t-SNE visualization of the VAE encoded features (p = 100) for all tumors.
Training
The current model training is explained in this notebook.
Tybalt dependencies are listed in environment.yml. To download
and activate this environment run:
# conda version 4.4.10
conda env create --force --file environment.yml
# activate environment
conda activate tybalt
Tybalt is also configured to train on GPUs using
gpu-environment.yml. To activate this environment run:
# conda version 4.4.10
conda env create --force --file gpu-environment.yml
# activate environment
conda activate tybalt-gpu
For a complete pipeline with reproducibility instructions, refer to run_pipeline.sh. Note that scripts originally written in Jupyter notebooks were ported to the scripts folder for pipeline purposes with:
jupyter nbconvert --to=script --FilesWriter.build_directory=scripts/nbconverted *.ipynb
Architecture
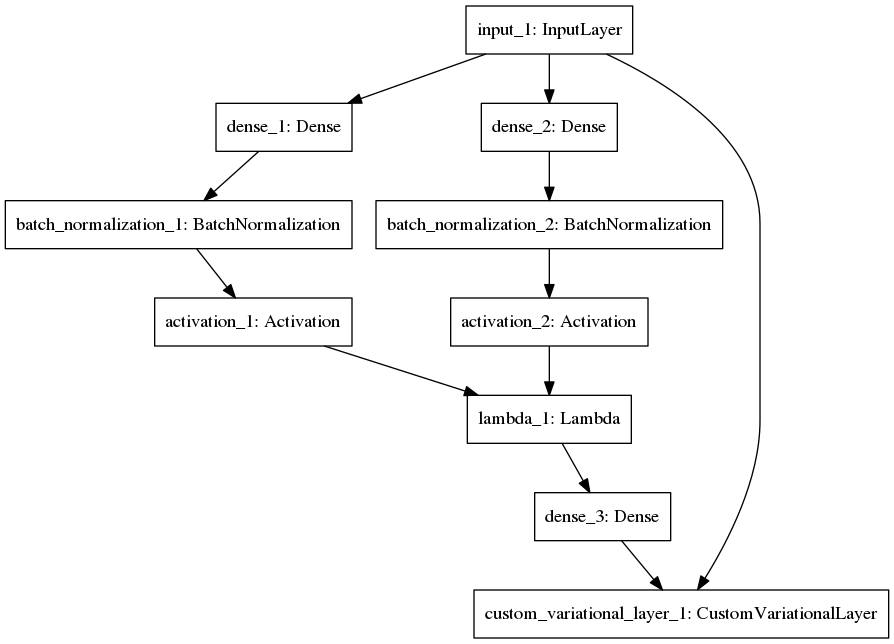
We select the top 5,000 most variably expressed genes by median absolute deviation. We compress this 5,000 vector of gene expression (for all samples) into two vectors of length 100; one representing the a mean and the other the variance. This vector can be sampled from to generate samples from an approximation of the data generating function. This hidden layer is then reconstructed back to the original dimensions. We use batch normalization and relu activation layers in the compression steps to prevent dead nodes and positive weights. We use a sigmoid activation in the decoder. We use the Keras library with a TensorFlow backend for training.
Parameter sweep
In order to select the most optimal parameters for the model, we ran a parameter search over a small grid of parameters. See parameter_sweep.md for more details.
Overall, we selected optimal learning rate = 0.0005, batch size = 50, and
epochs = 100. Because training did not improve much between 50 and 100 epochs,
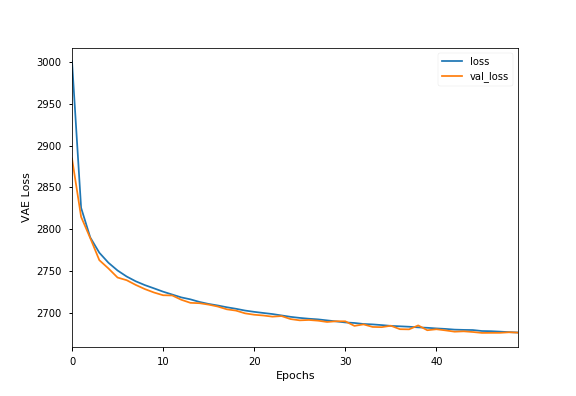
we used a 50 epoch model. Training and validation loss across 50 training epochs
for the optimal model is shown below.
Model Evaluation
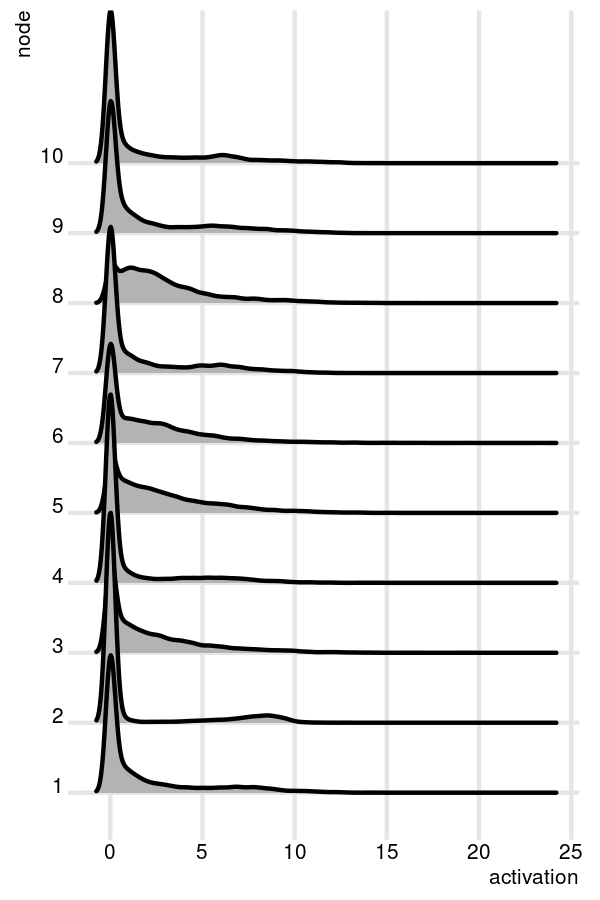
After training with optimal hyper parameters, the unsupervised model can be interpreted. For example, we can observe the distribution of activation patterns for all tumors across specific nodes. The first 10 nodes (of 100) are visualized below.
In this scenario, each node activation pattern contributes uniquely to each tumor and may represent specific gene expression signatures of biological significance. The distribution is heavily right skewed, with some nodes capturing slightly bimodal attributes.