ctrdata for aggregating and analysing clinical trials
The package ctrdata provides functions for retrieving (downloading)
information on clinical trials from public registers, and for
aggregating and analysing this information; it can be used for the
- EU Clinical Trials Register (“EUCTR”, https://www.clinicaltrialsregister.eu/)
- ClinicalTrials.gov (“CTGOV”, https://clinicaltrials.gov/)
- ISRCTN (https://www.isrctn.com/)
The motivation is to understand trends in design and conduct of trials,
their availability for patients and their detailled results. ctrdata
is a package for the R system, but other
systems and tools can be used with the databases created by it. This
README was reviewed on 2022-11-20 for version 1.11.1.
Main features:
-
Protocol-related trial information is easily retrieved (downloaded): Users define a query in a register’s web interface and then use
ctrdatato retrieve in one step all trials resulting from the query. Results-related trial information and personal annotations can be including during retrieval. Synonyms of an active substance can also be found. -
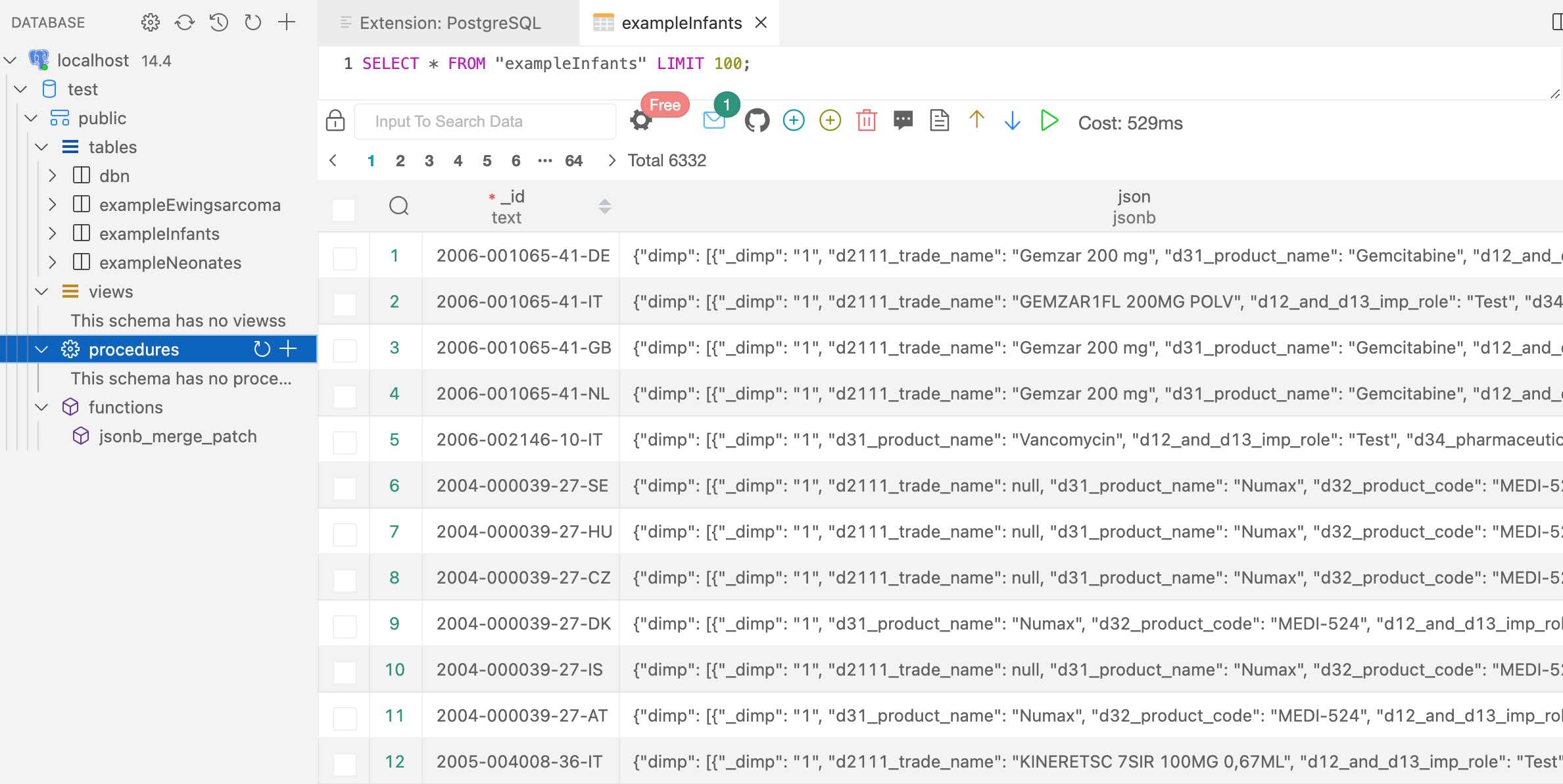
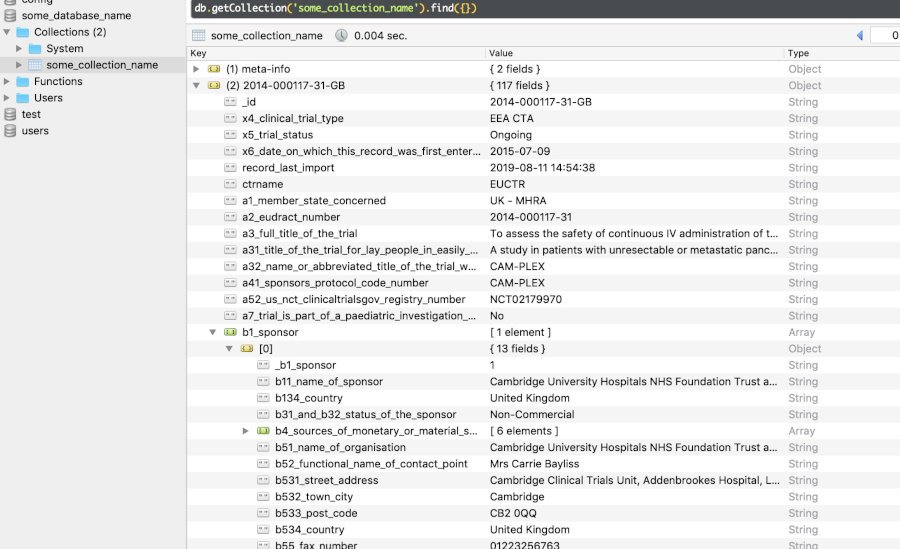
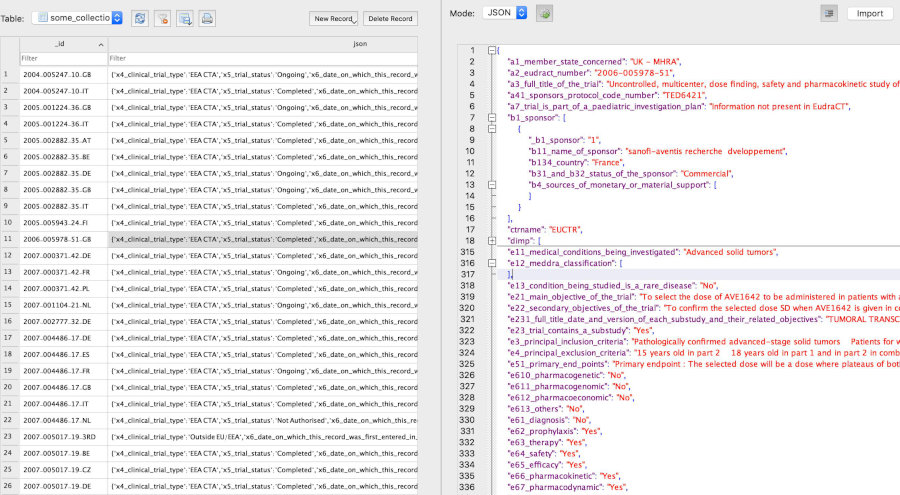
Retrieved (downloaded) trial information is transformed and stored in a document-centric database, for fast and offline access. Uses
DuckDB(🔔 new in version 1.11.0),PostgreSQL(new in version 1.9.0),RSQLiteorMongoDBas databases, via R packagenodbi: see section Databases below. Easily re-run a previous query to update a database. -
Analysis can be done with
R(usingctrdataconvenience functions) or others systems. Unique (de-duplicated) trial records are identified across registers.ctrdatacan merge and recode information (fields) and also provides easy access even to deeply-nested fields (new in version 1.4.0).
Remember to respect the registers’ terms and conditions (see
ctrOpenSearchPagesInBrowser(copyright = TRUE)). Please cite this
package in any publication as follows: “Ralf Herold (2022). ctrdata:
Retrieve and Analyze Clinical Trials in Public Registers. R package
version 1.11.0, https://cran.r-project.org/package=ctrdata”.
Installation
1. Install package ctrdata in R
Package ctrdata is on
CRAN and on
GitHub. Within
R, use the following commands to install
package ctrdata:
# Install CRAN version:
install.packages("ctrdata")
# Alternatively, install development version:
install.packages("devtools")
devtools::install_github("rfhb/ctrdata", build_vignettes = TRUE)These commands also install the package dependencies, which are nodbi,
jsonlite, httr, curl, clipr, xml2, rvest and stringi.
2. Command line tools perl, sed and php (5.2 or higher)
These are required for ctrLoadQueryIntoDb(), the main function of
package ctrdata (see Example workflow); the
function also checks if the tools can be used.
-
For MS Windows, install
Cygwin: InR, runctrdata::installCygwinWindowsDoInstall()for an automated minimal installation. Alternatively, manually install Cygwin with packagesperl,php-jsoncandphp-simplexmlintoc:\cygwin. The installation needs about 160 MB disk space; no administrator credentials needed. -
In macOS including 11 Big Sur, these are already installed; as alternative and
🔔 for macOS 12 Monterey and above,homebrewcan be used:brew install php, which seems to include the libraries required forctrdata. -
In Linux, these are usually already installed; tools to install vary by distribution (e.g.,
sudo apt install php-cli php-xml php-json).
Overview of functions in ctrdata
The functions are listed in the approximate order of use in a user’s workflow (in bold, main functions). See also the package documentation overview.
| Function name | Function purpose |
|---|---|
ctrOpenSearchPagesInBrowser() |
Open search pages of registers or execute search in web browser |
ctrFindActiveSubstanceSynonyms() |
Find synonyms and alternative names for an active substance |
ctrGetQueryUrl() |
Import from clipboard the URL of a search in one of the registers |
ctrLoadQueryIntoDb() |
Retrieve (download) or update, and annotate, information on trials from a register and store in a collection in a database |
dbQueryHistory() |
Show the history of queries that were downloaded into the collection |
dbFindIdsUniqueTrials() |
Get the identifiers of de-duplicated trials in the collection |
dbFindFields() |
Find names of variables (fields) in the collection |
dbGetFieldsIntoDf() |
Create a data frame (or tibble) from trial records in the database with the specified fields |
dfTrials2Long() |
Transform the data.frame from dbGetFieldsIntoDf() into a long name-value data.frame, including deeply nested fields |
dfName2Value() |
From a long name-value data.frame, extract values for variables (fields) of interest (e.g., endpoints) |
dfMergeTwoVariablesRelevel() |
Merge two simple variables into a new variable, optionally map values to a new set of values |
installCygwinWindowsDoInstall() |
Convenience function to install a Cygwin environment (MS Windows only) |
If package dplyr is loaded, a tibble is returned instead of a
data.frame.
Databases that can be used with ctrdata
Package ctrdata retrieves trial information and stores it in a
database collection, which has to be given as a connection object to
parameter con for several ctrdata functions; this connection object
is created in slightly different ways for the three supported database
backends that can be used with ctrdata as shown in the table. For a
speed comparison, see the nodbi
documentation.
Besides ctrdata functions below, any such a connection object can
equally be used with functions of other packages, such as nodbi (last
row in table) or, in case of MongoDB as database backend, mongolite
(see vignettes).
| Purpose | Function call |
|---|---|
| Create SQLite database connection | dbc <- nodbi::src_sqlite(dbname = "name_of_my_database", collection = "name_of_my_collection") |
| Create MongoDB database connection | dbc <- nodbi::src_mongo(db = "name_of_my_database", collection = "name_of_my_collection") |
| Create PostgreSQL database connection | dbc <- nodbi::src_postgres(dbname = "name_of_my_database"); dbc[["collection"]] <- "name_of_my_collection" |
| Create DuckDB database connection | dbc <- nodbi::src_duckdb(dbname = "name_of_my_database", collection = "name_of_my_collection") |
Use connection with ctrdata functions |
ctrdata::{ctrLoadQueryIntoDb, dbQueryHistory, dbFindIdsUniqueTrials, dbFindFields, dbGetFieldsIntoDf}(con = dbc, ...) |
Use connection with nodbi functions |
e.g., nodbi::docdb_query(src = dbc, key = dbc$collection, ...) |
Vignettes
- Install R package ctrdata
- Retrieve clinical trial information
- Summarise and analyse clinical trial information
Example workflow
The aim is to download protocol-related trial information and tabulate the trials’ status of conduct.
- Attach package
ctrdata:
library(ctrdata)- See help to get started with
ctrdata:
help("ctrdata-package")- Information on trial registers that can be used with
ctrdata:
help("ctrdata-registers")- Open registers’ advanced search pages in browser:
ctrOpenSearchPagesInBrowser()
# Please review and respect register copyrights:
ctrOpenSearchPagesInBrowser(copyright = TRUE)-
Adjust search parameters and execute search in browser
-
When trials of interest are listed in browser, copy the address from the browser’s address bar to the clipboard
-
Search used in this example: https://www.clinicaltrialsregister.eu/ctr-search/search?query=cancer&age=under-18&phase=phase-one&status=completed
-
Get address from clipboard:
q <- ctrGetQueryUrl()
# * Using clipboard content as register query URL: https://www.clinicaltrialsregister.eu/ctr-search/search?query=cancer&age=under-18&phase=phase-one&status=completed
# * Found search query from EUCTR: query=cancer&age=under-18&phase=phase-one&status=completed
q
# query-term query-register
# 1 query=cancer&age=under-18&phase=phase-one&status=completed EUCTR- Retrieve protocol-related information, transform and save to database:
The database collection is specified first, using nodbi (see above for
how to specify PostgreSQL, RSQlite, DuckDB or MongoDB as
backend); then, trial information is retrieved and loaded into the
collection:
# Connect to (or newly create) an SQLite database
# that is stored in a file on the local system:
db <- nodbi::src_sqlite(
dbname = "some_database_name.sqlite_file",
collection = "some_collection_name"
)
# See section Databases below
# for MongoDB as alternative
# Retrieve trials from public register:
ctrLoadQueryIntoDb(
queryterm = q,
con = db
)
# * Found search query from EUCTR: query=cancer&age=under-18&phase=phase-one&status=completed
# (1/3) Checking trials in EUCTR:
# Retrieved overview, multiple records of 66 trial(s) from 4 page(s) to be downloaded
# Checking helper binaries: done
# Downloading trials (4 pages in parallel)...
# Note: register server cannot compress data, transfer takes longer, about 0.4s per trial
# Pages: 4 done, 0 ongoing
# (2/3) Converting to JSON, 248 records converted
# (3/3) Importing JSON records into database...
# = Imported or updated 248 records on 66 trial(s)
# * Updated history ("meta-info" in "some_collection_name")Under the hood, scripts euctr2json.sh and xml2json.php (in
ctrdata/exec) transform EUCTR plain text files and CTGOV as well as
ISRCTN XML files to ndjson format, which is imported into the
database collection.
- Analyse
Tabulate the status of trials that are part of an agreed paediatric development program (paediatric investigation plan, PIP):
# Get all records that have values in the fields of interest:
result <- dbGetFieldsIntoDf(
fields = c(
"a7_trial_is_part_of_a_paediatric_investigation_plan",
"p_end_of_trial_status",
"a2_eudract_number"
),
con = db
)
# Find unique trial identifiers for trials that have nore than
# one record, for example for several EU Member States:
uniqueids <- dbFindIdsUniqueTrials(con = db)
# Searching for duplicate trials...
# - Getting trial ids, 279 found in collection
# - Finding duplicates among registers' and sponsor ids...
# - 208 EUCTR _id were not preferred EU Member State record for 71 trials
# - Keeping 71 records from EUCTR
# = Returning keys (_id) of 71 records in collection "some_collection_name"
# Keep only unique / de-duplicated records:
result <- subset(
result,
subset = `_id` %in% uniqueids
)
# Tabulate the selected clinical trial information:
with(
result,
table(
p_end_of_trial_status,
a7_trial_is_part_of_a_paediatric_investigation_plan
)
)
# a7_trial_is_part_of_a_paediatric_investigation_plan
# p_end_of_trial_status Information not present in EudraCT No Yes
# Completed 6 32 16
# GB - no longer in EU/EEA 0 7 5
# Ongoing 0 1 0
# Prematurely Ended 1 2 0
# Restarted 0 1 0-
Add records from another register (CTGOV) into the same collection
-
Search used in this example: https://clinicaltrials.gov/ct2/results?cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug
# Retrieve trials from another register:
ctrLoadQueryIntoDb(
queryterm = "cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug",
register = "CTGOV",
con = db
)
# * Found search query from CTGOV: cond=neuroblastoma&rslt=With&recrs=e&age=0&intr=Drug
# (1/3) Checking trials in CTGOV:
# Retrieved overview, records of 44 trial(s) are to be downloaded
# Checking helper binaries: done
# Downloading: 620 kB
# (2/3) Converting to JSON, 44 records converted
# (3/3) Importing JSON records into database...
# = Imported or updated 43 trial(s)
# * Updated history ("meta-info" in "some_collection_name")-
Add records from a third register (ISRCTN) into the same collection
-
Search used in this example: https://www.isrctn.com/search?q=neuroblastoma
# Retrieve trials from another register:
ctrLoadQueryIntoDb(
queryterm = "https://www.isrctn.com/search?q=neuroblastoma",
con = db
)
# * Found search query from ISRCTN: q=neuroblastoma
# (1/3) Checking trials in ISRCTN:
# Retrieved overview, records of 9 trial(s) are to be downloaded
# Checking helper binaries: done
# Downloading: 90 kB
# (2/3) Converting to JSON, 9 records converted
# (3/3) Importing JSON records into database...
# = Imported or updated 9 trial(s)
# * Updated history ("meta-info" in "some_collection_name")- Result-related trial information
Analyse some simple result details (see vignette for more examples):
# Get all records that have values in any of the specified fields
result <- dbGetFieldsIntoDf(
fields = c(
"clinical_results.baseline.analyzed_list.analyzed.count_list.count",
"clinical_results.baseline.group_list.group",
"clinical_results.baseline.analyzed_list.analyzed.units",
"study_design_info.allocation",
"location"
),
con = db
)
# Transform all fields into long name - value format
result <- dfTrials2Long(df = result)
# Total 6386 rows, 12 unique names of variables
# [1.] get counts of subjects for all arms into data frame
# This count is in the group where either its title or its
# description starts with "Total"
nsubj <- dfName2Value(
df = result,
valuename = "clinical_results.baseline.analyzed_list.analyzed.count_list.count.value",
wherename = paste0(
"clinical_results.baseline.group_list.group.title|",
"clinical_results.baseline.group_list.group.description"
),
wherevalue = "^Total"
)
# [2.] count number of sites
nsite <- dfName2Value(
df = result,
# some ctgov records use
# location.name, others use
# location.facility.name
valuename = "^location.*name$"
)
# count
nsite <- tapply(
X = nsite[["value"]],
INDEX = nsite[["_id"]],
FUN = length,
simplify = TRUE
)
nsite <- data.frame(
"_id" = names(nsite),
nsite,
check.names = FALSE,
stringsAsFactors = FALSE,
row.names = NULL
)
# [3.] randomised?
ncon <- dfName2Value(
df = result,
valuename = "study_design_info.allocation"
)
# merge sets
nset <- merge(nsubj, nsite, by = "_id")
nset <- merge(nset, ncon, by = "_id")
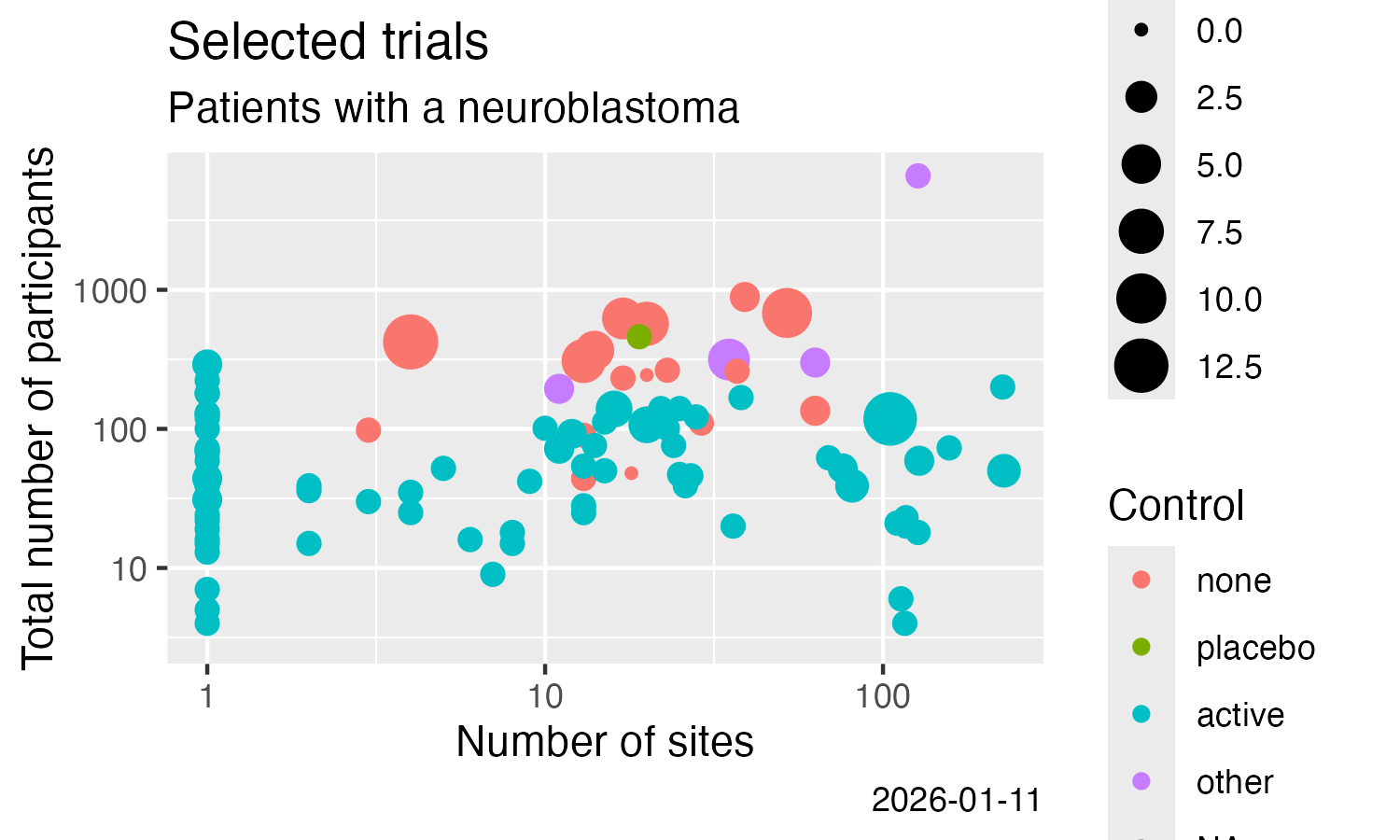
# Example plot
library(ggplot2)
ggplot(data = nset) +
labs(
title = "Neuroblastoma trials with results",
subtitle = "clinicaltrials.gov"
) +
geom_point(
mapping = aes(
x = nsite,
y = value.x,
colour = value.y == "Randomized"
)
) +
scale_x_log10() +
scale_y_log10() +
xlab("Number of sites") +
ylab("Total number of subjects") +
labs(colour = "Randomised?")
ggsave(
filename = "man/figures/README-ctrdata_results_neuroblastoma.png",
width = 5, height = 3, units = "in"
)- Retrieve protocols, statistical analysis plans and other documents
into a local folder
./files/
# eudract files are downloaded as part of results
ctrLoadQueryIntoDb(
queryterm = q,
euctrresults = TRUE,
euctrresultspdfpath = "./files/",
con = db
)
# ctgov files can separately be downloaded
sapply(
unlist(strsplit(
dbGetFieldsIntoDf(
fields = "provided_document_section.provided_document.document_url",
con = db
)[[2]],
split = " / "
)),
function(f) download.file(f, paste0("./files/", gsub("[/:]", "-", f)))
)Meta
Package ctrdata has been used for: Blogging on Innovation coming to
paediatric
research
and a Report on The impact of collaboration: The value of UK medical
research to EU science and
health
Additional features under consideration
-
Retrieve previous versions of protocol- or results-related information. The challenge is that, apparently, initial versions cannot be queried and historical versions can only be retrieved one-by-one and not in structured format.
-
Merge results-related fields retrieved from different registers (e.g., corresponding endpoints). The challenge is the incomplete congruency and different structure of fields.
Acknowledgements
-
Data providers and curators of the clinical trial registers. Please review and respect their copyrights and terms and conditions, see
ctrOpenSearchPagesInBrowser(copyright = TRUE). -
Package
ctrdatahas been made possible building on the work done for R, curl, httr, xml2, rvest, mongolite, jsonlite, nodbi, RPostgres, RSQLite and clipr.
Issues and notes
-
Please file issues and bugs here. Also check out how to handle some of the closed issues, e.g. on C stack usage too close to the limit and on a SSL certificate problem: unable to get local issuer certificate
-
Information in trial registers may not be fully correct; see for example this publication on CTGOV.
-
No attempts were made to harmonise field names between registers (nevertheless,
dfMergeTwoVariablesRelevel()can be used to merge and map two variables / fields into one).