easystats / Parameters
Programming Languages
Projects that are alternatives of or similar to Parameters
parameters 
Describe and understand your model’s parameters!
parameters’ primary goal is to provide utilities for processing the parameters of various statistical models (see here for a list of supported models). Beyond computing p-values, CIs, Bayesian indices and other measures for a wide variety of models, this package implements features like bootstrapping of parameters and models, feature reduction (feature extraction and variable selection), or tools for data reduction like functions to perform cluster, factor or principal component analysis.
Another important goal of the parameters package is to facilitate and streamline the process of reporting results of statistical models, which includes the easy and intuitive calculation of standardized estimates or robust standard errors and p-values. parameters therefor offers a simple and unified syntax to process a large variety of (model) objects from many different packages.
Installation
Run the following to install the stable release of parameters from CRAN:
install.packages("parameters")
Or this one to install the latest development version:
install.packages("remotes")
remotes::install_github("easystats/parameters")
Documentation
Click on the buttons above to access the package documentation and the easystats blog, and check-out these vignettes:
- Summary of Model Parameters
- Standardized Model Parameters
- Robust Estimation of Standard Errors, Confidence Intervals and p-values
- Model Parameters and Missing Data
- Feature reduction (PCA, cMDS, ICA…)
- Structural models (EFA, CFA, SEM…)
- Parameters selection
- A Practical Guide for Panel Data Analysis
Contributing and Support
In case you want to file an issue or contribute in another way to the package, please follow this guide. For questions about the functionality, you may either contact us via email or also file an issue.
Features
Model’s parameters description
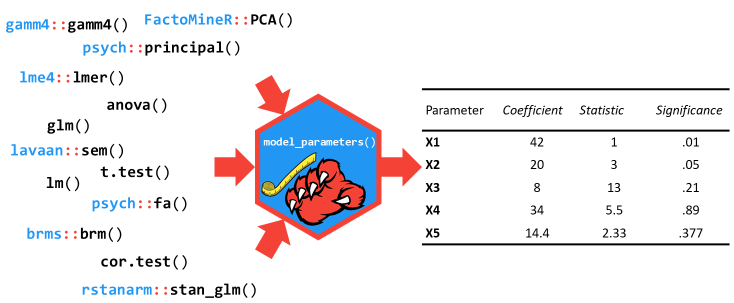
The
model_parameters()
function (that can be accessed via the parameters() shortcut) allows
you to extract the parameters and their characteristics from various
models in a consistent way. It can be considered as a lightweight
alternative to broom::tidy(),
with some notable differences:
- The column names of the returned data frame are specific to their
content. For instance, the column containing the statistic is named
following the statistic name, i.e., t, z, etc., instead of a
generic name such as statistic (however, you can get standardized
(generic) column names using
standardize_names()). - It is able to compute or extract indices not available by default, such as p-values, CIs, etc.
- It includes feature engineering capabilities, including parameters bootstrapping.
Classical Regression Models
model <- lm(Sepal.Width ~ Petal.Length * Species + Petal.Width, data = iris)
# regular model parameters
model_parameters(model)
#> Parameter | Coefficient | SE | 95% CI | t(143) | p
#> -------------------------------------------------------------------------------------------
#> (Intercept) | 2.89 | 0.36 | [ 2.18, 3.60] | 8.01 | < .001
#> Petal.Length | 0.26 | 0.25 | [-0.22, 0.75] | 1.07 | 0.287
#> Species [versicolor] | -1.66 | 0.53 | [-2.71, -0.62] | -3.14 | 0.002
#> Species [virginica] | -1.92 | 0.59 | [-3.08, -0.76] | -3.28 | 0.001
#> Petal.Width | 0.62 | 0.14 | [ 0.34, 0.89] | 4.41 | < .001
#> Petal.Length * Species [versicolor] | -0.09 | 0.26 | [-0.61, 0.42] | -0.36 | 0.721
#> Petal.Length * Species [virginica] | -0.13 | 0.26 | [-0.64, 0.38] | -0.50 | 0.618
# standardized parameters
model_parameters(model, standardize = "refit")
#> Parameter | Coefficient | SE | 95% CI | t(143) | p
#> -------------------------------------------------------------------------------------------
#> (Intercept) | 3.59 | 1.30 | [ 1.01, 6.17] | 2.75 | 0.007
#> Petal.Length | 1.07 | 1.00 | [-0.91, 3.04] | 1.07 | 0.287
#> Species [versicolor] | -4.62 | 1.31 | [-7.21, -2.03] | -3.53 | < .001
#> Species [virginica] | -5.51 | 1.38 | [-8.23, -2.79] | -4.00 | < .001
#> Petal.Width | 1.08 | 0.24 | [ 0.59, 1.56] | 4.41 | < .001
#> Petal.Length * Species [versicolor] | -0.38 | 1.06 | [-2.48, 1.72] | -0.36 | 0.721
#> Petal.Length * Species [virginica] | -0.52 | 1.04 | [-2.58, 1.54] | -0.50 | 0.618
Mixed Models
library(lme4)
model <- lmer(Sepal.Width ~ Petal.Length + (1|Species), data = iris)
# model parameters with CI, df and p-values based on Wald approximation
model_parameters(model)
#> Parameter | Coefficient | SE | 95% CI | t(146) | p
#> ------------------------------------------------------------------
#> (Intercept) | 2.00 | 0.56 | [0.90, 3.10] | 3.56 | < .001
#> Petal.Length | 0.28 | 0.06 | [0.17, 0.40] | 4.75 | < .001
# model parameters with CI, df and p-values based on Kenward-Roger approximation
model_parameters(model, df_method = "kenward")
#> Parameter | Coefficient | SE | 95% CI | t | df | p
#> -------------------------------------------------------------------------
#> (Intercept) | 2.00 | 0.57 | [0.07, 3.93] | 3.53 | 2.67 | 0.046
#> Petal.Length | 0.28 | 0.06 | [0.16, 0.40] | 4.58 | 140.98 | < .001
Structural Models
Besides many types of regression models and packages, it also works for other types of models, such as structural models (EFA, CFA, SEM…).
library(psych)
model <- psych::fa(attitude, nfactors = 3)
model_parameters(model)
#> # Rotated loadings from Factor Analysis (oblimin-rotation)
#>
#> Variable | MR1 | MR2 | MR3 | Complexity | Uniqueness
#> ------------------------------------------------------------
#> rating | 0.90 | -0.07 | -0.05 | 1.02 | 0.23
#> complaints | 0.97 | -0.06 | 0.04 | 1.01 | 0.10
#> privileges | 0.44 | 0.25 | -0.05 | 1.64 | 0.65
#> learning | 0.47 | 0.54 | -0.28 | 2.51 | 0.24
#> raises | 0.55 | 0.43 | 0.25 | 2.35 | 0.23
#> critical | 0.16 | 0.17 | 0.48 | 1.46 | 0.67
#> advance | -0.11 | 0.91 | 0.07 | 1.04 | 0.22
#>
#> The 3 latent factors (oblimin rotation) accounted for 66.60% of the total variance of the original data (MR1 = 38.19%, MR2 = 22.69%, MR3 = 5.72%).
Variable and parameters selection
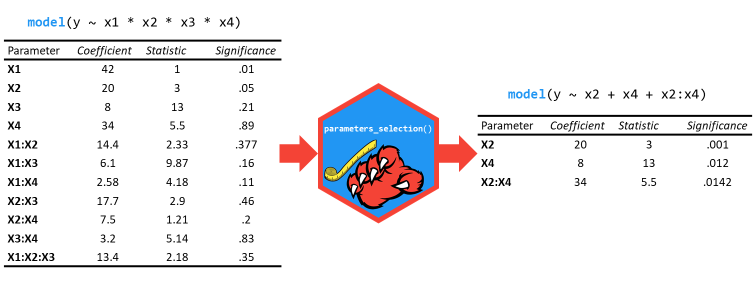
select_parameters()
can help you quickly select and retain the most relevant predictors
using methods tailored for the model type.
library(dplyr)
lm(disp ~ ., data = mtcars) %>%
select_parameters() %>%
model_parameters()
#> Parameter | Coefficient | SE | 95% CI | t(26) | p
#> -----------------------------------------------------------------------
#> (Intercept) | 141.70 | 125.67 | [-116.62, 400.02] | 1.13 | 0.270
#> cyl | 13.14 | 7.90 | [ -3.10, 29.38] | 1.66 | 0.108
#> hp | 0.63 | 0.20 | [ 0.22, 1.03] | 3.18 | 0.004
#> wt | 80.45 | 12.22 | [ 55.33, 105.57] | 6.58 | < .001
#> qsec | -14.68 | 6.14 | [ -27.31, -2.05] | -2.39 | 0.024
#> carb | -28.75 | 5.60 | [ -40.28, -17.23] | -5.13 | < .001
Miscellaneous
This packages also contains a lot of other useful functions:
Describe a Distribution
data(iris)
describe_distribution(iris)
#> Variable | Mean | SD | IQR | Range | Skewness | Kurtosis | n | n_Missing
#> ----------------------------------------------------------------------------------------
#> Sepal.Length | 5.84 | 0.83 | 1.30 | [4.30, 7.90] | 0.31 | -0.55 | 150 | 0
#> Sepal.Width | 3.06 | 0.44 | 0.52 | [2.00, 4.40] | 0.32 | 0.23 | 150 | 0
#> Petal.Length | 3.76 | 1.77 | 3.52 | [1.00, 6.90] | -0.27 | -1.40 | 150 | 0
#> Petal.Width | 1.20 | 0.76 | 1.50 | [0.10, 2.50] | -0.10 | -1.34 | 150 | 0
Citation
In order to cite this package, please use the following citation:
- Lüdecke D, Ben-Shachar M, Patil I, Makowski D (2020). parameters: Extracting, Computing and Exploring the Parameters of Statistical Models using R. Journal of Open Source Software, 5(53), 2445. doi: 10.21105/joss.02445
Corresponding BibTeX entry:
@Article{,
title = {parameters: Extracting, Computing and Exploring the Parameters of Statistical Models using {R}.},
volume = {5},
doi = {10.21105/joss.02445},
number = {53},
journal = {Journal of Open Source Software},
author = {Daniel Lüdecke and Mattan S. Ben-Shachar and Indrajeet Patil and Dominique Makowski},
year = {2020},
pages = {2445},
}