MouseLand / Suite2p
Projects that are alternatives of or similar to Suite2p
suite2p 
Pipeline for processing two-photon calcium imaging data.
Copyright (C) 2018 Howard Hughes Medical Institute Janelia Research Campus
suite2p includes the following modules:
- Registration
- Cell detection
- Spike detection
- Visualization GUI
This code was written by Carsen Stringer and Marius Pachitariu.
For support, please open an issue.
The reference paper is here.
The deconvolution algorithm is based on this paper, with settings based on this paper.
See this twitter thread for GUI demonstrations.
The matlab version is available here. Note that the algorithm is older and will not work as well on non-circular ROIs.
Lectures on how suite2p works are available here.
Read the Documentation at https://suite2p.readthedocs.io/
Installation
Install an Anaconda distribution of Python -- Choose Python 3.x and your operating system. Note you might need to use an anaconda prompt if you did not add anaconda to the path.
- Download the
environment.ymlfile from the repository. You can do this by cloning the repository, or copy-pasting the text from the file into a text document on your local computer. - Open an anaconda prompt / command prompt with
condafor python 3 in the path - Change directories to where the
environment.ymlis and runconda env create -f environment.yml - Activate the environment with
conda activate suite2p - Now run
python -m suite2pand you're all set. - Running the command
suite2p --versionin the terminal will print the install version of suite2p.
If you have an older suite2p environment you can remove it with conda env remove -n suite2p before creating a new one.
Note you will always have to run conda activate suite2p before you run suite2p. Conda ensures mkl_fft and numba run correctly and quickly on your machine. If you want to run jupyter notebooks in this environment, then also conda install jupyter.
To upgrade the suite2p (package here), run the following in the environment:
pip install --upgrade suite2p
Common issues
If you are on Yosemite Mac OS, PyQt doesn't work, and you won't be able to install suite2p. More recent versions of Mac OS are fine.
The software has been heavily tested on Windows 10 and Ubuntu 18.04, and less well tested on Mac OS. Please post an issue if you have installation problems. The registration step runs faster on Ubuntu than Windows, so if you have a choice we recommend using the Ubuntu OS.
Installation of github version for developers
- Clone the repository and
cd suite2pin an anaconda prompt / command prompt withcondafor python 3 in the path - Run
conda env create -f environment.yml - To activate this new environment, run
conda activate suite2p(you will have to activate every time you want to run suite2p) - Install the local version of suite2p into this environment in develop mode, with everything to get test data:
pip install -e .[data] - Download the data with
dvc pull -r gdrive-travis(read-only access). If you need to have read/write access to the files, rundvc pulland follow the authentication instructions in the output. Note: do not commit the json credentials file that comes out afterwards, git should ignore it. - Run tests:
python setup.py test
Examples
An example dataset is provided here. It's a single-plane, single-channel recording.
Getting started
The quickest way to start is to open the GUI from a command line terminal. You might need to open an anaconda prompt if you did not add anaconda to the path. Make sure to run this from a directory in which you have WRITE access (suite2p saves a couple temporary files in your current directory):
suite2p
Then:
- File -> Run suite2p (or ctrl+r)
- Setup a configuration
- -> Add directory which contains tiffs to data_path (can be multiple folders, but add them one at a time)
- -> OR choose an h5 file which has a key with the data, data shape should be time x pixels x pixels (you can type in the key name for the data after you choose the file)
- -> Add save_path ((otherwise the data directory is used as save path))
- -> Add fast_disk (this is where the binary file of registered data will be created, choose an SSD for this path) ((otherwise the save path is used as the fast disk path))
- Set some parameters (see full list below). At the minimum:
nplanes, nchannels, tau, fs
- Press run and wait. Messages should start appearing in the embedded command line.
- When the run is finished, the results will open in the GUI window and there you can visualize and refine the results (see below).
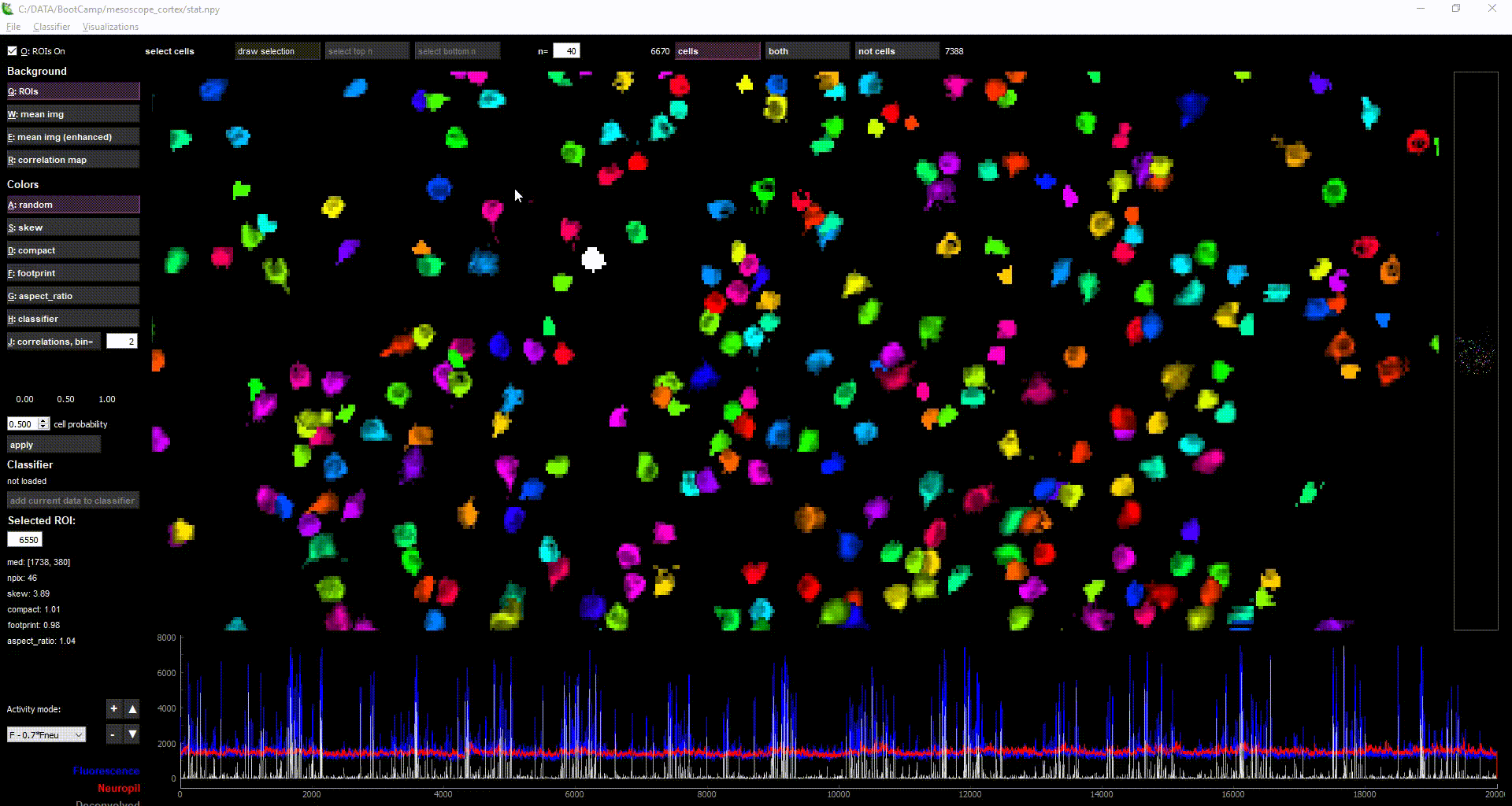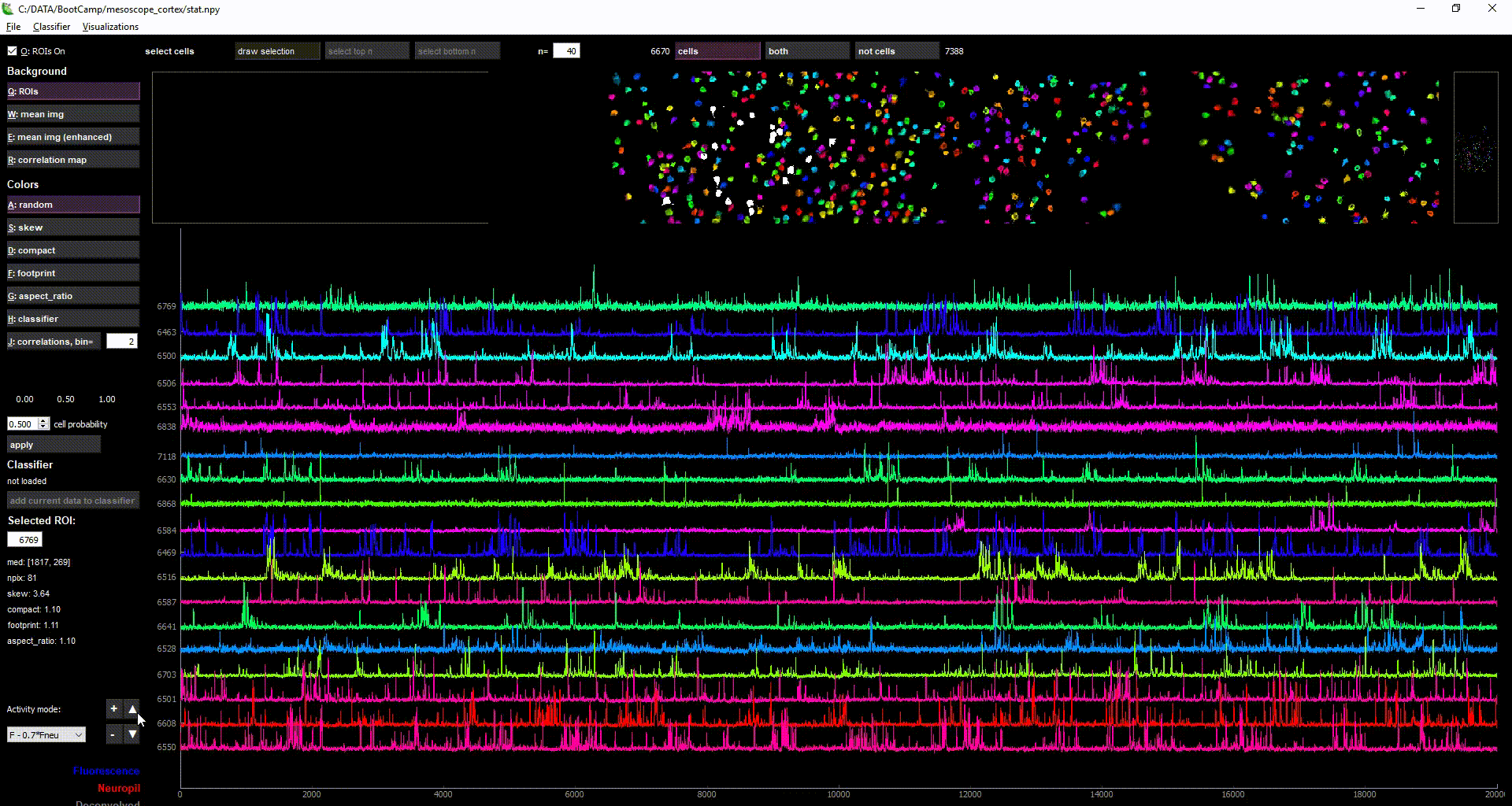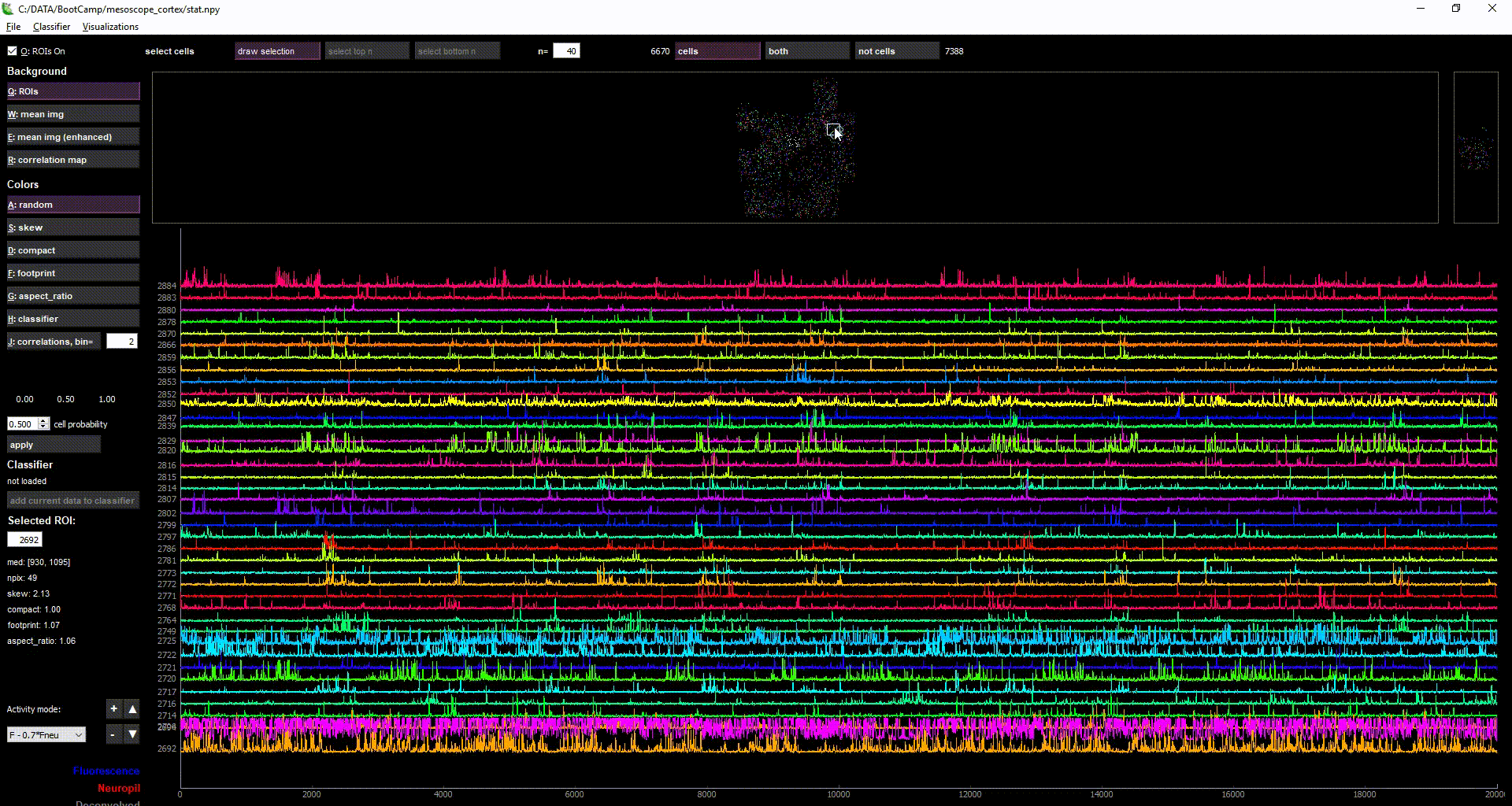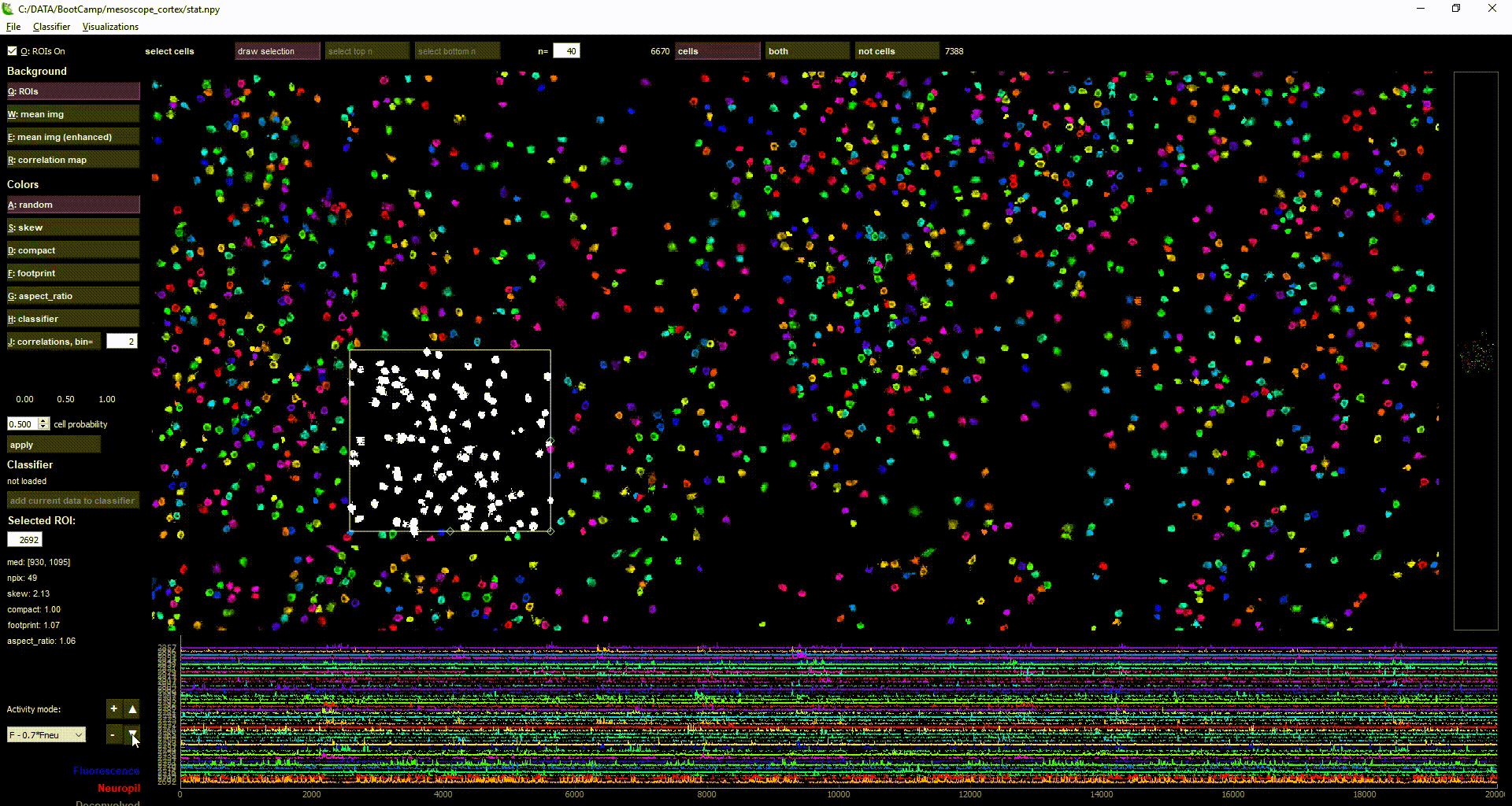
Using the GUI
suite2p output goes to a folder called "suite2p" inside your save_path, which by default is the same as the data_path. If you ran suite2p in the GUI, it loads the results automatically. Otherwise, load the results with File -> Load results.
The GUI serves two main functions:
- Checking the quality of the data and results.
- there are currently several views such as the enhanced mean image, the ROI masks, the correlation map, the correlation among cells, and the ROI+neuropil traces
- by selecting multiple cells (with "Draw selection" or ctrl+left-click), you can view the activity of multiple ROIs simultaneously in the lower plot
- there are also population-level visualizations, such as rastermap
- Classify ROIs into cell / not cell (left and right views respectively)
- the default classifier included should work well in a variety of scenarios.
- a user-classifier can be learnt from manual curation, thus adapting to the statistics of your own data.
- the GUI automatically saves which ROIs are good in "iscell.npy". The second column contains the probability that the ROI is a cell based on the currently loaded classifier.
Main GUI controls (works in all views):
- Pan = Left-Click + drag
- Zoom = (Scroll wheel) OR (Right-Click + drag)
- Full view = Double left-click OR escape key
- Swap cell = Right-click on the cell
- Select multiple cells = (Ctrl + left-click) OR (SHIFT + left-click) AND/OR ("Draw selection" button)
You can add your manual curation to a pre-built classifier by clicking "Add current data to classifier". Or you can make a brand-new classifier from a list of "iscell.npy" files that you've manually curated. The default classifier in the GUI is initialized as the suite2p classifier, but you can overwrite it by adding to it, or loading a different classifier and saving it as the default. The default classifier is used in the pipeline to produce the initial "iscell.npy" file.
Other ways to call Suite2p
- From the command line:
suite2p --ops <path to ops.npy> --db <path to db.npy>
- From Python/Jupyter
from suite2p.run_s2p import run_s2p
ops1 = run_s2p(ops, db)
See our example jupyter notebook here. It also explains how to batch-run suite2p.
Outputs
F.npy: array of fluorescence traces (ROIs by timepoints)
Fneu.npy: array of neuropil fluorescence traces (ROIs by timepoints)
spks.npy: array of deconvolved traces (ROIs by timepoints)
stat.npy: array of statistics computed for each cell (ROIs by 1)
ops.npy: options and intermediate outputs
iscell.npy: specifies whether an ROI is a cell, first column is 0/1, and second column is probability that the ROI is a cell based on the default classifier
Dependencies
suite2p relies on the following excellent packages (which are automatically installed with conda/pip if missing):
- rastermap
- pyqtgraph
- PyQt5
- numpy (>=1.16.0)
- numba
- mkl_fft
- scanimage-tiff-reader
- scipy
- h5py
- scikit-learn
- scanimage-tiff-reader
- tifffile
- natsort
- matplotlib (not for plotting (only using hsv_to_rgb and colormap function), should not conflict with PyQt5)
Logo
Logo was designed by Shelby Stringer and Chris Czaja.