peace195 / Multitask Learning Protein Prediction
Multitask learning: protein secondary structure prediction, b-values prediction and solvent-accessibility prediction
Programming Languages
python 139335 projects - #7 most used programming language
Projects that are alternatives of or similar to Multitask Learning Protein Prediction
Ner LstmNamed Entity Recognition using multilayered bidirectional LSTM
Stars: ✭ 532 (+2700%)
Mutual labels: lstm
Conv EmotionThis repo contains implementation of different architectures for emotion recognition in conversations.
Stars: ✭ 646 (+3300%)
Mutual labels: lstm
Python autocompleteA simple neural network for python autocompletion
Stars: ✭ 779 (+4000%)
Mutual labels: lstm
VadVoice activity detection (VAD) toolkit including DNN, bDNN, LSTM and ACAM based VAD. We also provide our directly recorded dataset.
Stars: ✭ 622 (+3173.68%)
Mutual labels: lstm
StockpricepredictionStock Price Prediction using Machine Learning Techniques
Stars: ✭ 700 (+3584.21%)
Mutual labels: lstm
Music2danceGenerating Dance steps for given music with deep learning
Stars: ✭ 17 (-10.53%)
Mutual labels: lstm
Ad examplesA collection of anomaly detection methods (iid/point-based, graph and time series) including active learning for anomaly detection/discovery, bayesian rule-mining, description for diversity/explanation/interpretability. Analysis of incorporating label feedback with ensemble and tree-based detectors. Includes adversarial attacks with Graph Convolutional Network.
Stars: ✭ 641 (+3273.68%)
Mutual labels: lstm
Seq2seq ChatbotChatbot in 200 lines of code using TensorLayer
Stars: ✭ 777 (+3989.47%)
Mutual labels: lstm
Video ClassificationTutorial for video classification/ action recognition using 3D CNN/ CNN+RNN on UCF101
Stars: ✭ 543 (+2757.89%)
Mutual labels: lstm
TelemanomA framework for using LSTMs to detect anomalies in multivariate time series data. Includes spacecraft anomaly data and experiments from the Mars Science Laboratory and SMAP missions.
Stars: ✭ 589 (+3000%)
Mutual labels: lstm
Deep Learning Time SeriesList of papers, code and experiments using deep learning for time series forecasting
Stars: ✭ 796 (+4089.47%)
Mutual labels: lstm
Qa使用深度学习算法实现的中文问答系统
Stars: ✭ 528 (+2678.95%)
Mutual labels: lstm
Text ClassificationImplementation of papers for text classification task on DBpedia
Stars: ✭ 682 (+3489.47%)
Mutual labels: lstm
Tensorflow OnnxConvert TensorFlow models to ONNX
Stars: ✭ 900 (+4636.84%)
Mutual labels: lstm
Tensorflow TutorialSome interesting TensorFlow tutorials for beginners.
Stars: ✭ 893 (+4600%)
Mutual labels: lstm
Getting Things Done With PytorchJupyter Notebook tutorials on solving real-world problems with Machine Learning & Deep Learning using PyTorch. Topics: Face detection with Detectron 2, Time Series anomaly detection with LSTM Autoencoders, Object Detection with YOLO v5, Build your first Neural Network, Time Series forecasting for Coronavirus daily cases, Sentiment Analysis with BERT.
Stars: ✭ 738 (+3784.21%)
Mutual labels: lstm
Descriptions
Multitask learning (secondary structure prediction, b-values prediction, solvent-accessibility prediction) can improve the prediction accuracy of protein secondary structure.
- We have to face with the class imbalance problem
- "foldername_cv": 5 fold cross validation
- Distribution of outputs:
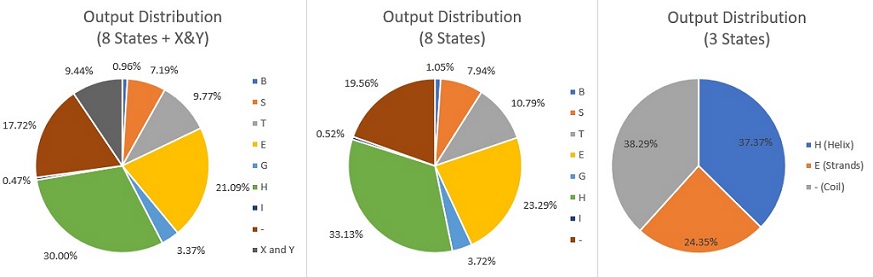
Data
The copyright belongs to http://rostlab.org/. It can not be public.
Data representation
Using Protvec (3-gram) and follow the vector addition rule. For example:
TNCDE = UTN + TNC + NCD + CDE + DEU
Multitask learning model
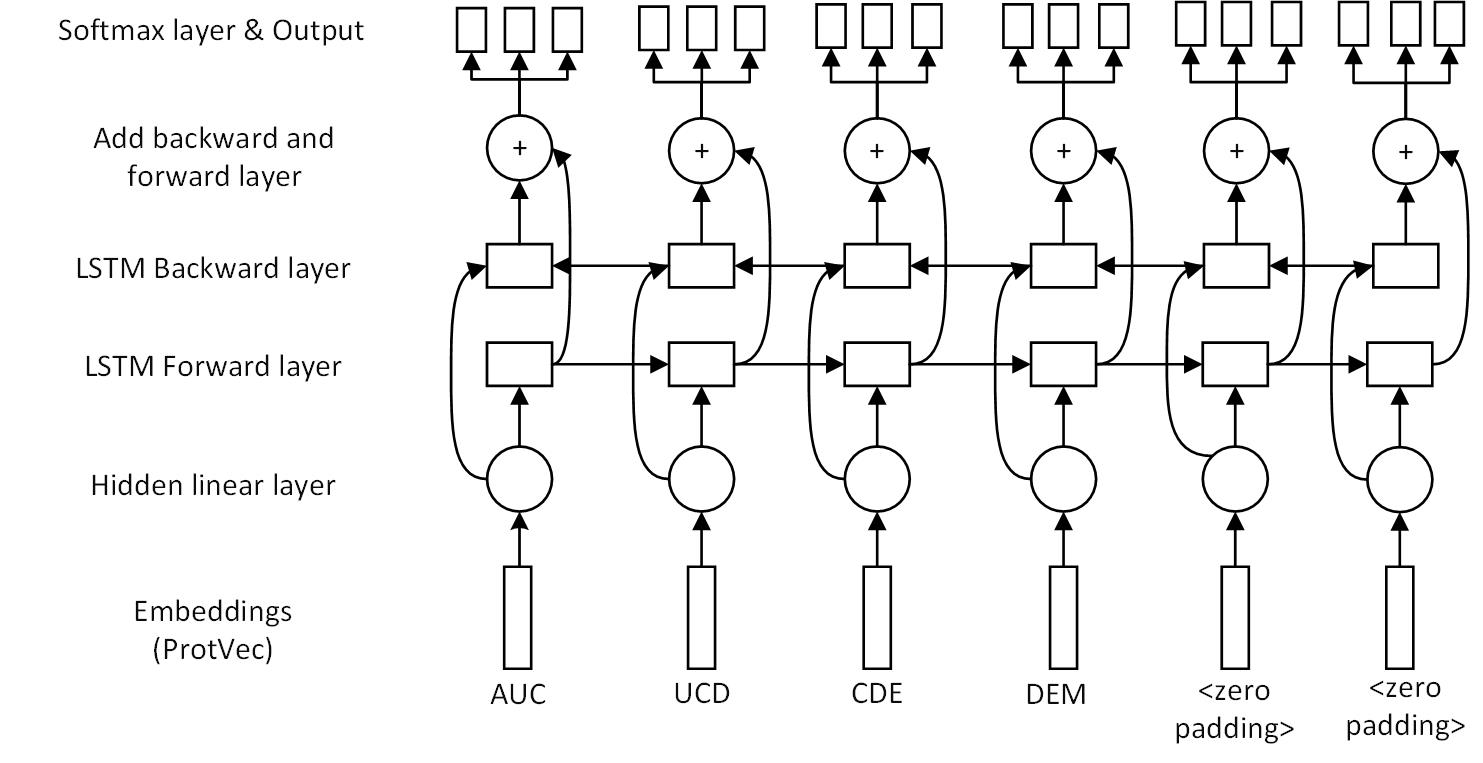
Results
3 states protein secondary structure)
Multi-task learning (3 tasks, 3 states):
-
Secondary Structure accuracy (3 states): 69.0%
-
Solvent Accessibility accuracy (3 states): 54.6%
-
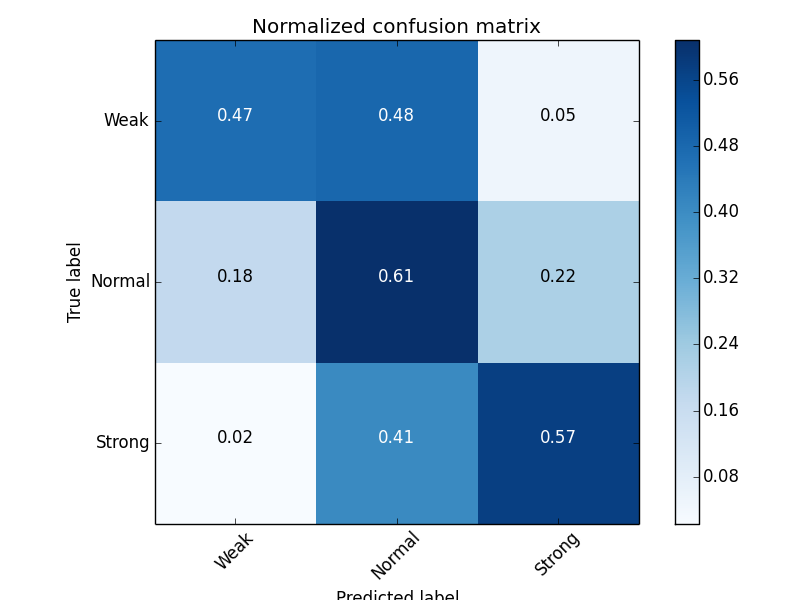
B-values accuracy (3 states): 59.1%
8 states protein secondary structure
Multi-task learning (3 tasks, 8 states):
-
Secondary Structure accuracy (8 states): 0.476
-
Solvent Accessibility accuracy (3 states): 0.548
-
B-values accuracy (3 states): 0.598
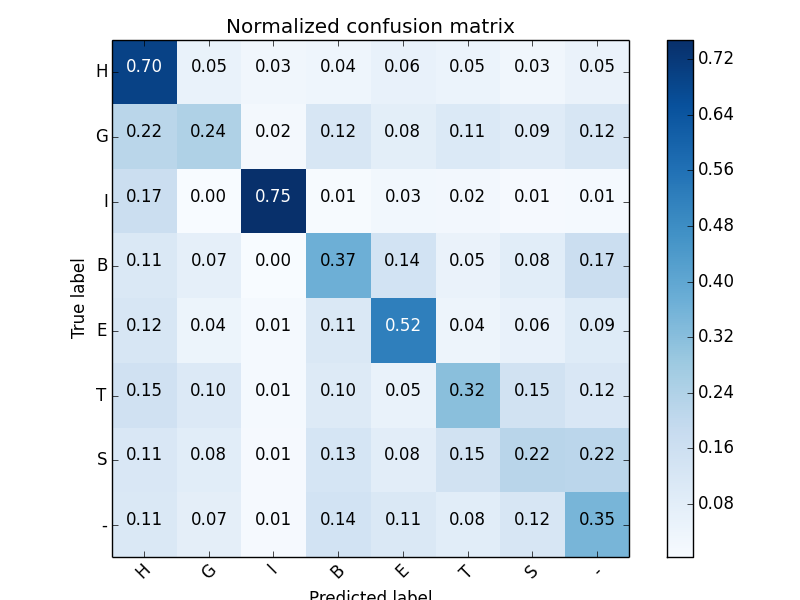
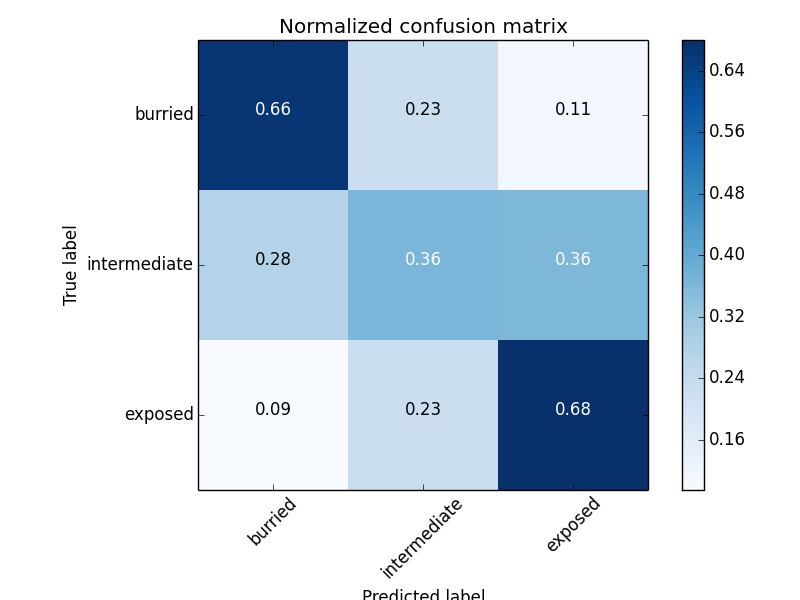
Prerequisites
- python 2.7
- tensorflow 1.4.0
- ProtVec
How to run
Go into each subfolder and run the code following:
Author
Binh Do
License
This project is licensed under the MIT License
Note that the project description data, including the texts, logos, images, and/or trademarks,
for each open source project belongs to its rightful owner.
If you wish to add or remove any projects, please contact us at
[email protected].





