ropensci / Restez
Programming Languages
Projects that are alternatives of or similar to Restez
Locally query GenBank 
NOTE:
restezis no longer available on CRAN due to the archiving of a key dependency. It can still be installed via GitHub. The issue is being dealt with and hopefully a new version ofrestezwill be available on CRAN soon.
Download parts of NCBI’s GenBank to a local folder and create a simple SQL-like database. Use ‘get’ tools to query the database by accession IDs. rentrez wrappers are available, so that if sequences are not available locally they can be searched for online through Entrez.
See the detailed tutorials for more information.
Introduction
Vous entrez, vous rentrez et, maintenant, vous …. restez!
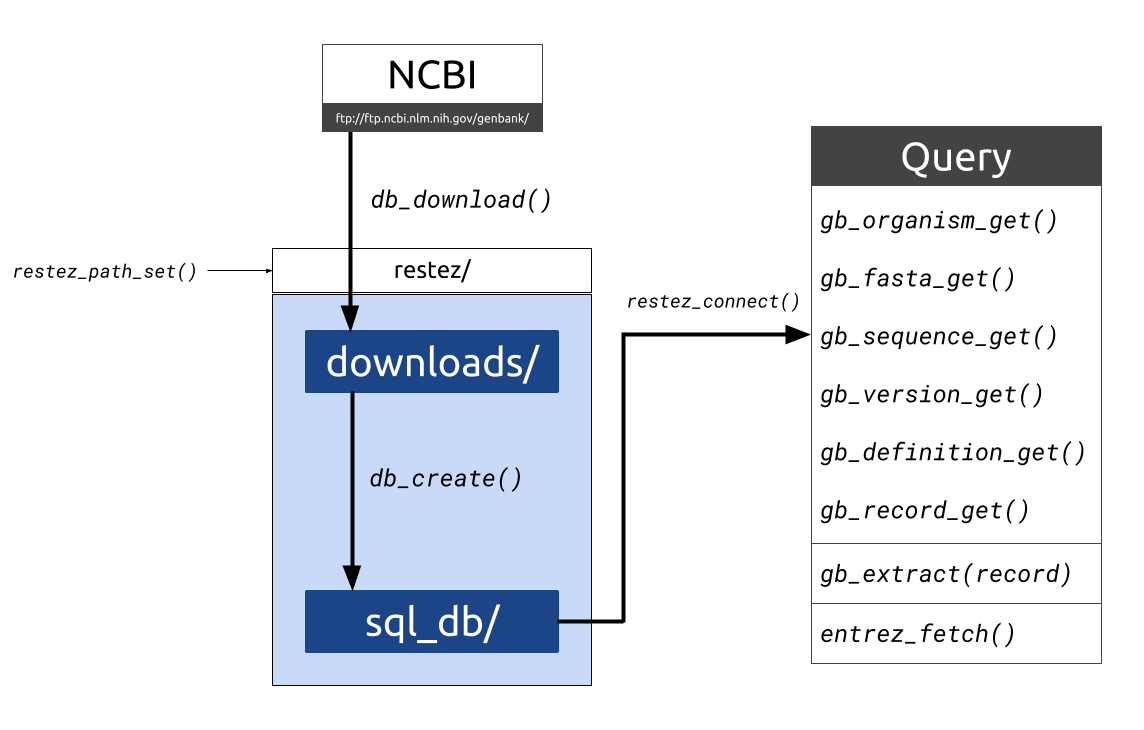
Downloading sequences and sequence information from GenBank and related NCBI taxonomic databases is often performed via the NCBI API, Entrez. Entrez, however, has a limit on the number of requests and downloading large amounts of sequence data in this way can be inefficient. For programmatic situations where multiple Entrez calls are made, downloading may take days, weeks or even months.
This package aims to make sequence retrieval more efficient by allowing a user to download large sections of the GenBank database to their local machine and query this local database either through package specific functions or Entrez wrappers. This process is more efficient as GenBank downloads are made via NCBI’s FTP using compressed sequence files. With a good internet connection and a middle-of-the-road computer, a database comprising 20 GB of sequence information can be generated in less than 10 minutes.
Installation
The package can currently only be installed through GitHub:
# install.packages("remotes")
remotes::install_github("ropensci/restez")
(It was previously available via CRAN but was archived due to a key dependency MonetDBLite being no longer available.)
Quick Examples
For more detailed information on the package’s functions and detailed guides on downloading, constructing and querying a database, see the detailed tutorials.
Setup
# Warning: running these examples may take a few minutes
library(restez)
#> -------------
#> restez v1.0.2
#> -------------
#> Remember to restez_path_set() and, then, restez_connect()
# choose a location to store GenBank files
restez_path_set(rstz_pth)
# Run the download function
db_download()
# after download, create the local database
db_create()
Query
# connect, ensure safe disconnect after finishing
restez_connect()
#> Remember to run `restez_disconnect()`
# get a random accession ID from the database
id <- sample(list_db_ids(), 1)
#> Warning in list_db_ids(): Number of ids returned was limited to [100].
#> Set `n=NULL` to return all ids.
# you can extract:
# sequences
seq <- gb_sequence_get(id)[[1]]
str(seq)
#> chr "ACTCTGACTTTTTACTGTATATAAAAACAGCTTTTTGGTTTATACTTGAATTCAGGAATAACCAAGCAGGTGTAAATATGCCAGCGCAAGAACAGCAAATTT"
# definitions
def <- gb_definition_get(id)[[1]]
print(def)
#> [1] "Unidentified RNA clone P10.7"
# organisms
org <- gb_organism_get(id)[[1]]
print(org)
#> [1] "unidentified"
# or whole records
rec <- gb_record_get(id)[[1]]
cat(rec)
#> LOCUS AF040899 102 bp RNA linear UNA 06-MAR-1998
#> DEFINITION Unidentified RNA clone P10.7.
#> ACCESSION AF040899
#> VERSION AF040899.1
#> KEYWORDS .
#> SOURCE unidentified
#> ORGANISM unidentified
#> unclassified sequences.
#> REFERENCE 1 (bases 1 to 102)
#> AUTHORS Pan,W.S., Ji,X.Y., Wang,H.T. and Zhong,Y.S.
#> TITLE RNA from plasma of patient NO.10
#> JOURNAL Unpublished
#> REFERENCE 2 (bases 1 to 102)
#> AUTHORS Pan,W.S., Ji,X.Y., Wang,H.T. and Zhong,Y.S.
#> TITLE Direct Submission
#> JOURNAL Submitted (31-DEC-1997) Department of Applied Molecular Biology,
#> Microbiology & Epidemiology Institution, 20 Dongdajie Street,
#> Fengtai, Beijing 100071, China
#> FEATURES Location/Qualifiers
#> source 1..102
#> /organism="unidentified"
#> /mol_type="genomic RNA"
#> /db_xref="taxon:32644"
#> /clone="P10.7"
#> /note="from the plasma of patient no.10, a person infected
#> by an unknown hepatitis virus"
#> ORIGIN
#> 1 actctgactt tttactgtat ataaaaacag ctttttggtt tatacttgaa ttcaggaata
#> 61 accaagcagg tgtaaatatg ccagcgcaag aacagcaaat tt
#> //
Entrez wrappers
# use the entrez_* wrappers to access GB data
res <- entrez_fetch(db = 'nucleotide', id = id, rettype = 'fasta')
cat(res)
#> >AF040899.1 Unidentified RNA clone P10.7
#> ACTCTGACTTTTTACTGTATATAAAAACAGCTTTTTGGTTTATACTTGAATTCAGGAATAACCAAGCAGG
#> TGTAAATATGCCAGCGCAAGAACAGCAAATTT
# if the id is not in the local database
# these wrappers will search online via the rentrez package
res <- entrez_fetch(db = 'nucleotide', id = c('S71333.1', id),
rettype = 'fasta')
#> [1] id(s) are unavailable locally, searching online.
cat(res)
#> >AF040899.1 Unidentified RNA clone P10.7
#> ACTCTGACTTTTTACTGTATATAAAAACAGCTTTTTGGTTTATACTTGAATTCAGGAATAACCAAGCAGG
#> TGTAAATATGCCAGCGCAAGAACAGCAAATTT
#>
#> >S71333.1 alpha 1,3 galactosyltransferase [New World monkeys, mermoset lymphoid cell line B95.8, mRNA Partial, 1131 nt]
#> ATGAATGTCAAAGGAAAAGTAATTCTGTCGATGCTGGTTGTCTCAACTGTGATTGTTGTGTTTTGGGAAT
#> ATATCAACAGCCCAGAAGGCTCTTTCTTGTGGATATATCACTCAAAGAACCCAGAAGTTGATGACAGCAG
#> TGCTCAGAAGGACTGGTGGTTTCCTGGCTGGTTTAACAATGGGATCCACAATTATCAACAAGAGGAAGAA
#> GACACAGACAAAGAAAAAGGAAGAGAGGAGGAACAAAAAAAGGAAGATGACACAACAGAGCTTCGGCTAT
#> GGGACTGGTTTAATCCAAAGAAACGCCCAGAGGTTATGACAGTGACCCAATGGAAGGCGCCGGTTGTGTG
#> GGAAGGCACTTACAACAAAGCCATCCTAGAAAATTATTATGCCAAACAGAAAATTACCGTGGGGTTGACG
#> GTTTTTGCTATTGGAAGATATATTGAGCATTACTTGGAGGAGTTCGTAACATCTGCTAATAGGTACTTCA
#> TGGTCGGCCACAAAGTCATATTTTATGTCATGGTGGATGATGTCTCCAAGGCGCCGTTTATAGAGCTGGG
#> TCCTCTGCGTTCCTTCAAAGTGTTTGAGGTCAAGCCAGAGAAGAGGTGGCAAGACATCAGCATGATGCGT
#> ATGAAGACCATCGGGGAGCACATCTTGGCCCACATCCAACACGAGGTTGACTTCCTCTTCTGCATGGATG
#> TGGACCAGGTCTTCCAAGACCATTTTGGGGTAGAGACCCTGGGCCAGTCGGTGGCTCAGCTACAGGCCTG
#> GTGGTACAAGGCAGATCCTGATGACTTTACCTATGAGAGGCGGAAAGAGTCGGCAGCATATATTCCATTT
#> GGCCAGGGGGATTTTTATTACCATGCAGCCATTTTTGGAGGAACACCGATTCAGGTTCTCAACATCACCC
#> AGGAGTGCTTTAAGGGAATCCTCCTGGACAAGAAAAATGACATAGAAGCCGAGTGGCATGATGAAAGCCA
#> CCTAAACAAGTATTTCCTTCTCAACAAACCCTCTAAAATCTTATCTCCAGAATACTGCTGGGATTATCAT
#> ATAGGCCTGCCTTCAGATATTAAAACTGTCAAGCTATCATGGCAAACAAAAGAGTATAATTTGGTTAGAA
#> AGAATGTCTGA
restez_disconnect()
Contributing
Want to contribute? Check the contributing page.
Version
Release version 1.
Licence
MIT
Citation
Bennett et al. (2018). restez: Create and Query a Local Copy of GenBank in R. Journal of Open Source Software, 3(31), 1102. https://doi.org/10.21105/joss.01102
References
Benson, D. A., Karsch-Mizrachi, I., Clark, K., Lipman, D. J., Ostell, J., & Sayers, E. W. (2012). GenBank. Nucleic Acids Research, 40(Database issue), D48–D53. http://doi.org/10.1093/nar/gkr1202
Winter DJ. (2017) rentrez: An R package for the NCBI eUtils API. PeerJ Preprints 5:e3179v2 https://doi.org/10.7287/peerj.preprints.3179v2

